CH 105 - Chemistry and
Society
Influenza: Overview
04/18/2008
How do you get influenza
Influenza is a highly communicable
respiratory virus. The picture below says it all.

credit: http://www.immunize.org/images/ca.d/ipcd1861/img0024.htm
| Clinical Features
(This section taken directly from:
http://virology-online.com/viruses/Influenza2.htm)
Following a typical incubation period of
48 hours, the typical symptoms of influenza appears. The onset is abrupt
with a marked fever, headache, photophobia, shivering, a dry cough,
malaise, myalgia, and a dry tickling throat. The fever is continuous and
lasts around 3 days. Influenza B infection is similar to influenza A,
but infection with influenza C is usually subclinical or very mild in
nature.
Complications
- 1. Tracheobronchitis and
bronchiolitis - A small proportion of
patients develop more sever respiratory symptoms where rales and
rhonchi are heard but the chest is radiologically clear. These
symptoms are more commonly seen in the elderly and patients with
COAD.
- 2. Pneumonia
- primary viral pneumonia or a secondary bacterial pneumonia may
develop. Primary viral pneumonia is relatively uncommon, but cases
have been demonstrated in many influenza epidemics. It may occur in
previously young and healthy persons, but are commonly associated
with patients with preexisting cadiovascular disease such as
Rheumatic fever. Secondary bacterial pneumonia is more common than
primary viral pneumonia. It was speculated that the high incidence
of deaths in young people during the Spanish influenza pandemics of
1917-1918 may have been due to secondary bacterial pneumonia in a
population generally debilitated by the effects the WWI.
- Secondary bacterial
pneumonia - usually occurs late in
the course of disease, after a period of improvement has been
observed for the acute disease. The symptoms and signs are that of a
typical bacterial pneumonia. S. aureus is most commonly
involved although S. pneumoniae and H. influenzae may
be found. There appears to be a good reason why S. aureus is
so commonly found in cases of secondary bacterial pneumonia.
Infection of cells by influenza A requires cleavage of the virus
haemagglutinin by proteases, and some strains of S. aureus
produces such enzymes. Thus S. aureus and influenza may
promote infection by the other. Influenza A by damage to the healthy
respiratory epithelium.
- Myositis and
myoglobinuria - In addition to myalgia,
which is characteristic of acute influenza infection, clinical
myositis and myoglobinuria may occur.
- Reye's syndrome
- Reye's syndrome is characterized by encephalopathy and fatty liver
degeneration. The disease has a 50% mortality amongst hospitalized
cases and had been associated with several viruses; such as
influenza A and B, Coxsackie B5, echovirus, HSV, VZV, CMV and
adenovirus.
- Other complications
- influenza infection have been implicated in acute viral
encephalitis and Guillain-Barre syndrome. Influenza A was also
associated with the cot death syndrome.
E. Laboratory Diagnosis
During epidemics, a presumptive
diagnosis can be made on the basis of the clinical symptoms. However,
influenza A and B can co-circulate, and mixed infections of influenza
and other viruses have been reported. Isolated cases of suspected
influenza should be investigated for these may represent the first cases
of an impending epidemic.
- Virus Isolation
- Throat swabs, NPA and nasal washings may be used for virus
isolation. It is reported that nasal washings are the best specimens
for virus isolation. The specimen may be inoculated in embryonated
eggs or tissue culture. 10-12 day embryonated eggs are used for
virus isolation. The specimen is inoculated into the amniotic
cavity. The virus replicates in the cells of the amniotic membrane
and large quantities are released back into the amniotic fluid.
After 2-3 days incubation, virus in the amniotic fluid can be
detected by adding aliquots of harvested amniotic fluid to chick,
guinea pig, or human erythrocytes. Pathological specimens can be
inoculated on to tissue cultures of kidney, chicks or a variety of
other species. Rhesus monkey cells are the most sensitive. Although
no CPE is produced, newly produced virus can be recognized by
haemadsorption using the cells in the tissue culture, and
haemagglutination using the culture medium which contains free virus
particles. Influenza B virus and occasionally influenza A will
produce a CPE in MDCK cells. Influenza viruses isolated from
embryonated eggs or tissue culture can be identified by serological
or molecular methods. Influenza viruses can be recognized as A, B,
or C types by the use of complement fixation tests against the
soluble antigen. (A soluble antigen is found for all influenza A, B
or C type virus but antibody against one type does not cross react
with the soluble antigen of the other. The further classification of
influenza isolates into subtypes and strains is a highly specialized
responsibility of the WHO reference laboratories. The HA type is
identified by HAI tests, the NA type is also identified.
- Rapid Diagnosis by
Immunofluorescence - cells from
pathological specimens may be examined for the presence of influenza
A and B antigens by indirect immunofluorescence. Although many
workers are convinced of the value of this technique, others have
been disappointed with the specificity of the antisera and the level
of background fluorescence that makes the test difficult to
interpret. EIA tests for the detection of influenza A viral antigens
are available that are easier to interpret than immunofluorescence.
PCR assays for the detection of influenza RNA have also been
developed but there usefulness in a clinical setting is highly
questionable.
- Serology
- Virus cannot be isolated from all cases of suspected infection.
More commonly, the diagnosis is made retrospectively by the
demonstration of a rise in serum antibody to the infecting virus.
CFT is the most common method used using the type specific soluble
antigen. However, the CF test is thought to have a low specificity.
A more specific test is the HAI test. Infection by influenza viruses
results in a rise in serum antibody titre, but the requirement for a
4-fold or greater rise in titre of HI of CF antibody reflects the
inaccuracy of these tests for detecting smaller increases in
antibody. A more precise method for measuring antibody is by SRH.
SRH is more sensitive than CF or HAI tests and has a greater degree
of precision. A 50% increase in zone area represents a rise in
antibody and is evidence of recent infection. Sera do not have to be
pretreated to remove non-specific inhibitors which plaque the HAI
test. SRH may well replace CF and HAI tests in diagnostic laboratory
in future.
|
Influenza Virus
The disease is caused by a virus.


credit:
http://www.microscopy.fsu.edu/cells/viruses/images/influenzafigure1.jpg credit: http://www.ou.edu/class/pheidole/bacteria.html
bacteriophage electronmicrograph
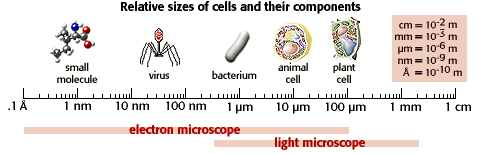
credit:
http://www.biology.arizona.edu/cell_bio/tutorials/cells/cells2.html
An electron micrograph picture of the virus in an
infected cell is shown below. The spherical-like particles are the
viruses.
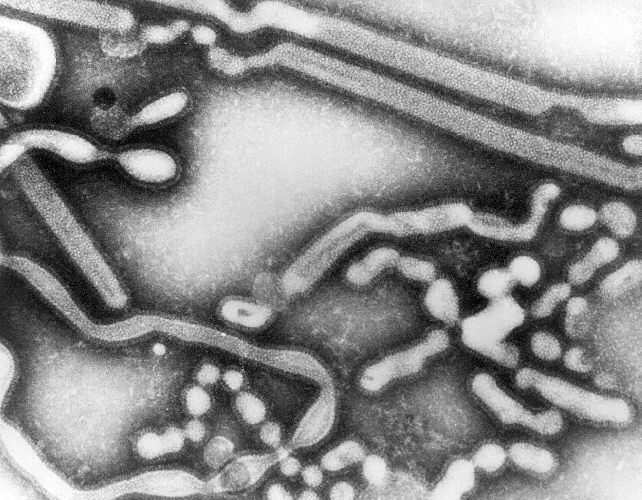
credit:
http://phil.cdc.gov/Phil/detail.asp?id=279
Life Cycle of the Virus

credit:
http://web.uct.ac.za/depts/mmi/jmoodie/flu2life.gif

credit:
http://www.mie.utoronto.ca/labs/lcdlab/biopic/fig/13.04.jpg
Virus Genome and Proteins
The genetic material in the influenza virus is RNA, not DNA as is found in
humans. Viruses, as opposed to bacteria and eukaryotic cells, can have
either DNA (single stranded or double stranded) or RNA (ss or ds) genomes.
The genetic material in most viruses consists of single ss or ds "molecule"
which contain many genes. Influenza is an example of a class of viruses
which contains several (7-8) different RNA molecules (or segments), not one long
RNA molecule. Replication of the RNA genome of influenza hence does not
require a DNA polymerase. Instead it contains an enzyme complex called a
transcriptase or RNA polymerase which replicates RNA into duplicate RNA molecules. the table below
shows the different RNA segments, their size (number of monomeric nucleotide
units) and the proteins produced when the RNA strands are translated into
proteins. The virus is a member of a class called Orthomyxoviridae.
These viruses have segmented ss-RNA genomes in which the RNA represents the
negative sense strand. This strand replicates when new viral is created.
The negative strand is also transcribed into a positive mRNA strand which is
then translated to produce viral proteins. (see life cycle above.) Both
replication and transcription of the RNA occur in the nucleus of the cell (very
unusual). Presumably the same enzyme (transcriptase/RNA polymerase) is
required for both activities.
| Segment: |
Size(nt) |
Polypeptide(s) |
Function |
| 1 |
2341 |
PB2 |
RNA Polymerase/Transcriptase: cap binding |
| 2 |
2341 |
PB1 |
RNA Polymerase/Transcriptase: elongation |
| 3 |
2233 |
PA |
RNA Polymerase/Transcriptase: protease activity (?) |
| 4 |
1778 |
HA |
Haemagglutinin |
| 5 |
1565 |
NP |
Nucleoprotein: RNA binding; part of transcriptase complex; nuclear/cytoplasmic
transport of vRNA |
| 6 |
1413 |
NA |
Neuraminidase: release of virus (N1, N2 in human viruses, 7 others
in other animals) |
| 7 |
1027 |
M1 |
Matrix protein: major component of virion |
| M2 * |
Integral membrane protein - ion channel |
| 8 |
890 |
NS1 |
Non-structural: nucleus; effects on cellular RNA transport,
splicing, translation |
| NS2 * |
Non-structural: nucleus+cytoplasm, function unknown |
* different reading frame;
credit:
http://www.tulane.edu/~dmsander/WWW/335/Orthomyxoviruses.html
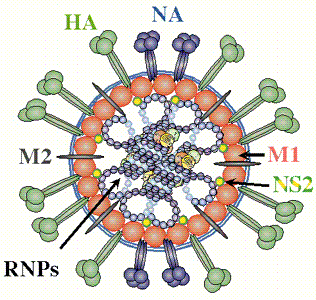
credit: � Paul Digard, Dept
Pathology, University of Cambridge
http://www-micro.msb.le.ac.uk/3035/Orthomyxoviruses.html
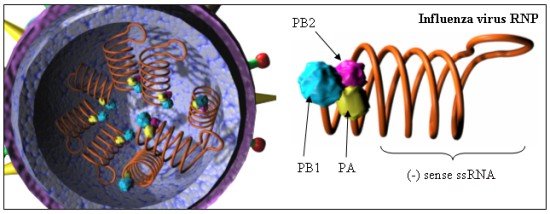
credit:
http://www.omedon.co.uk/influenza/influenza/
Additional information on the genome and
proteins of influenza virus can be found at the National Center for
Biotechnology Information (NCBI) links below:
There are 3 types of flu viruses: A, B, and C.
Two (A and B) cause human influenzas. Type A can infect many species of animals,including poulty, swine, horses, humans,
etc. Aquatic birds are the natural reservoir for the virus, which infects
their gut without causing
illness.
The two major
surfaces proteins of the virus are hemagglutinin (HA) and neuraminidase (NA). HA
is responsible for virus
binding
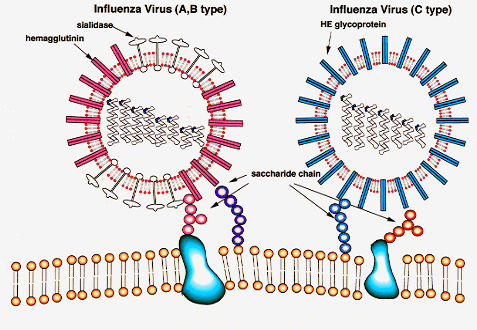
Credit:
http://www.gifu-u.ac.jp/~kassei/influenza%20virus.gif
Hemagglutinin (HA)
Hemagglutinin (HA) responsible for:
- virus binding to host receptor
- internalization of virus
- membrane fusion of internalized virus endosome
(intracellular vesicle with virus)
- Animation:
Influenza entering cell
Hemagglutinin:
- most abundant protein on viral surface (as surmised by antibody
formation)
- 15 avian and mammalian variants have been identified (based on
antibody studies); only 3 adapt to humans in last 100 yr, giving
pandemic strains H1 (1918), H2 (957) and H3 (1968)
- 3 recent avian variants (H5, H7, and H9) jump directly to humans recently;
but low human to human transmissibility.
- What if avian strain jump with high human transmissibility?
Structure HA:
- mature form is homotrimer (3 identical
protein subunits), MW 220,000; multiple sites
for covalent attachment of sugars. Hemagglutinin is a glycoprotein.
- each monomer synthesized as single polypeptide chain precursor
(HA0) that
is cleaved into HA1 and HA2 subunits bythe protease (a protein enzyme)
trypsin in epithelial
cells of lung.
- structure known for human (H3), swine (H9), avian (H5) subtypes.
 Chime Model:
Hemagglutinin antigen |
1918 Active Form Jmol
(Discusison)
Chime Model:
Hemagglutinin antigen |
1918 Active Form Jmol
(Discusison)
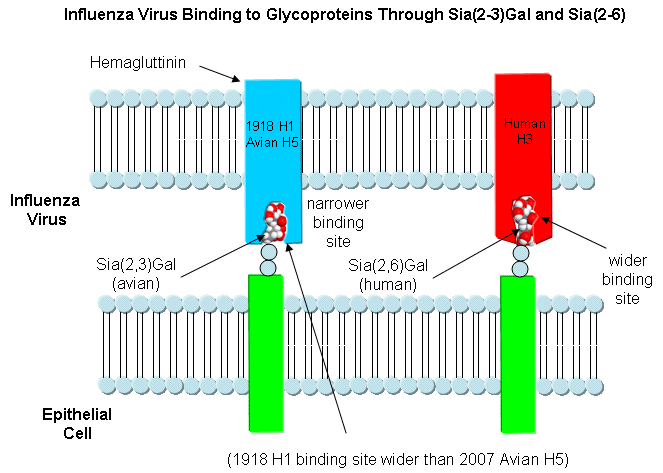
HA binding site:
- bind to a specific sugar, sialic acid
(Sia), which is covalently attached to many cell membrane glycoproteins
- sialic acid usually connected through
an α(2,3) or
α(2,6)
link to another sugar, galactose (Gal) on cell surface glycoproteins which
contain carbohydrates bound to a specific amino acid, asparagine.
- subtypes found in avian (and equine) influenza
isolates bind preferentially to
Sia (α2,3)
jGal
which predominates in avian GI tract where viruses replicate
- Human influenza isolates prefer Sia α(2,6)
Gal. Human virus of H1, H2, and H3 subtype (cause 1918,
1957, and 1968
pandemics) recognize Sia α(2,6)
Gal, major form in
human respiratory tract.
- swine influenza HA bind to Sia
α(2,6)
Gal and some Sia
(α2,3)
Gal both of which found in swine.
This suggests the swine as an intermediary genetic "mixing vessel".
|
Sia
α(2,6)
Gal (Human) |
Sia
α(2,3)
Gal (Avian and some Swine) |
|
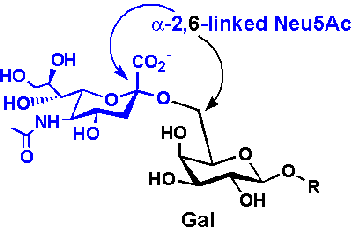 |
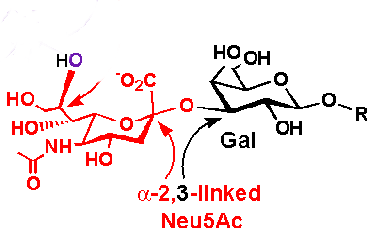 |
|
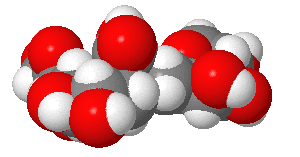
(made with
Sweet,
with an OH, not AcNH on sialic acid on C5) |
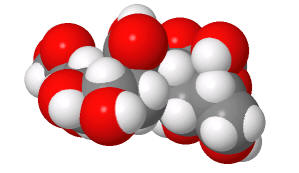
(made with
Sweet,
with an OH, not AcNH on sialic acid on C5) |
Structures from: http://www.bme.jhu.edu/~kjyarema/CellSurCarbo1.htm#linkages
Links:
For cross-species transfer
need changes in binding specificity.
- for human virus H2 and H3, need minimum 2 changes in receptor
binding site: Asparagine (polar) 226 to Leucine (nonpolar), and Glycine
(nonpolar, small) 228 to Serine (polar) to shift from avian to human receptor binding.
(Amino acid structure)
- for human H1, can bind to human receptor w/o change in
asparagine 226 and glycine 228.
- What the is basis for H1 binding to human receptor?
Neuraminidase (NA)
Activity: The virus before it leaves the
cell forms a bud on the intracellular side of the cell with the HA and NA in the
cell membrane of the host cell. The virus in this state would not leave
the cell since its HA molecules would interact with sialic acid residues in the
host cell membrane, holding the virus in the membrane. Neuraminidase
cleaves (hydrolysis) from cell surface glycoproteins, allowing the virus to
complete the budding process and be released from the cell as new viruses.
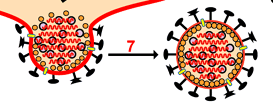
credit:
http://www.ncbi.nlm.nih.gov/genomes/VIRUSES/virusreplication_scheme.html
 Chime Model:
Neuraminidase
another model
|
Chime Model:
Neuraminidase
another model
|
Links:











![]() Chime Model:
Hemagglutinin antigen |
1918 Active Form Jmol
(Discusison)
Chime Model:
Hemagglutinin antigen |
1918 Active Form Jmol
(Discusison)

![]() Chime Model:
Neuraminidase
another model
|
Chime Model:
Neuraminidase
another model
|