Biochemistry Online: An Approach Based on Chemical Logic

CHAPTER 5 - BINDING
D: BINDING AND THE
CONTROL OF GENE
TRANSCRIPTION
BIOCHEMISTRY - DR. JAKUBOWSKI
Last Updated: 03/30/16
|
Learning Goals/Objectives for Chapter 5D: After class and this reading, students will be able to
|
D6. Studying the Interactome
Yeast Two Hybrid System
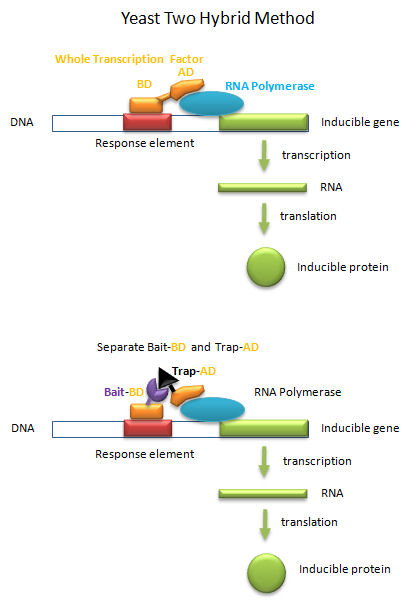
Transcription factors must do more than bind to upstream targets (response elements) on DNA. They must also interact with and activate the transcription machinery, including RNA polymerase and other assembled proteins, for transcription to occur. To accomplish both tasks, protein transcription factors usually consist of two distinct and often separable domains, a DNA Binding Domain (DNA-BD) and an Activation Domain (AD). The BD usually contained the DNA binding motifs discussed above. In a clever feat of genetic manipulation, scientist have taken the part of the transcription factor gene encoding the DNA-BD and fused it to a gene for a protein called the bait protein. Likewise the other part of the transcription factor gene encoding the AD is fused it to a gene for another protein, the target, that could bind the bait protein. Plasmids with the gene constructs are added to yeast. By themselves, the separated BD and AD can not activate transcription from a gene in the yeast that is inducible by the whole transcription factor. However, if the BD-Bait gene and the AD-Target genes are both added on separate plasmids, and both fusion genes are ultimately transcribed and the fusion RNA translated into fusions proteins, then transcription from the inducible gene can occur if the bait and target part domains of the fusion proteins bind to each other, allowing binding of the BD domain and the AD domain to their target sites, leading to gene transcription. This yeast two-hybrid method has allowed the determination of protein binding partners, part of what is now termed the interactome.
Figure: Yeast Two-Hybrid
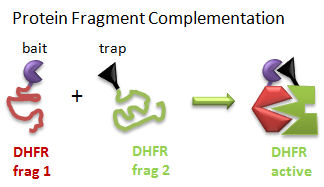
Protein Fragment Complementation (PFC) (Tarassov, K. et al): A gene encoding one fragment of a reporter enzyme, dihydrofolate reductase (DHFR), is fused to a gene for a bait protein and inserted into a plasmid. A second gene representing the second fragment of a reporter enzyme is linked to possible target protein genes. In a cell transformed with both plasmids, the reporter gene will ultimately display enzymatic activity only if the translated bait protein binds to a translated target protein, allowing the two fragments of the enzyme to interact and fold collectively into a holo-, active enzyme activity. The reporter gene was a mutant of DHFR that was resistant to an inhibitor, methotrexate. Functional DHFR activity leads to cell growth in the presence of methotrexate.
Figure: Protein Fragment Complementation
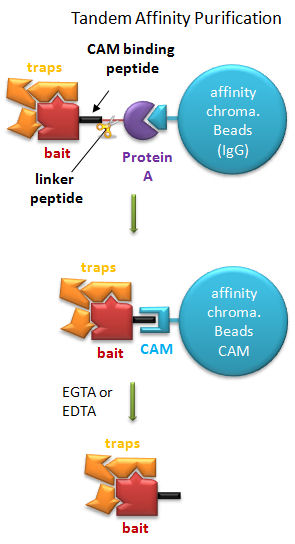
Tandem Affinity Purification (TAP) (Rigaut, G. et al) A generic protein purification method for protein complex characterization and proteome exploration. Nature Biotechnology 17. 1030 (1999): A gene for a bait protein is linked sequentially to genes encoding two separate tags, one for Protein A (which binds immunoglobulin G - IgG) and a calmodulin binding peptide (which binds the protein calmodulin in a process which requires calcium. In between the genes for the two tags is a nucleotide sequence encoding a short, protease sensitive linker peptide. The gene construct is introduced into yeast and expression induced. Cell are lysed and the extract applied to affinity chromatography beads containing covalently attached IgG, which binds the Protein A tag. After extensive washing the bound bait protein, with associated target proteins, is eluted by proteolysis of the peptide linker. The eluate is applied to a second affinity column containing covalently attached calmodulin. After washing, the trap and associated target proteins are eluted with a calcium chelator (EGTA or EDTA) as the interaction of calmodulin with the second tag, calmodulin binding peptide, requires calcium. Eluted target proteins can be identified by 2D PAGE or mass spectrometry.
Figure: Tandem Affinity Purification
A recent comparison of the Y2H, PFC, and TAP methods was made (Jensen, L. and Bork, 2008; Yu, H et al, 2008). Not unexpectedly, the Y2H was best at determining nuclear protein interactions, the TAP for cytoplasmic and abundant proteins, and PFC for transmembrane proteins. All suffer somewhat in identifying transient protein complexes.
Navigation
Return to Chapter 5D: Binding and the Control of Gene Transcription
Return to Biochemistry Online Table of Contents
Archived version of full Chapter 5D: Binding and the Control of Gene Transcription

Biochemistry Online by Henry Jakubowski is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.