04/17/16
Biochemistry Online: An Approach Based on Chemical Logic

The Origin of Life
K. Fixing CO2
Of the five different pathways known to fix CO2, all require ATP except one. That one is present in both methanogen, which produce methane from CO2 and H2 and in the acetogens, which produce acetate (CH3CO2-) in the form of acetylCoA. The simpler reactions of forming acetic acid and methane are shown below:
2CO2 + 4H2 --> CH3CO2H + 2H2O.
CO2 + 4H2 --> CH4 + 2H2O.
The DG0 values for these reaction (calculated using DG0f for gas phase H2, CO2 and CH4 and liquid acetic acid and water are -75 and -131 kJ, respectively, at 250C, showing that they are thermodynamically favored. Making AcetylCoA, a "high" energy molecule compared to its hydrolysis products (as is ATP) from acetic acid and CoASH, a would require energy input. A proton gradient is the likely source.
Some bacteria and Achaea cells (primordial or present) use the reductive acetylCoA pathway, also known as the Wood-Ljungdahl pathway, to form, in a noncyclic process, acetyl CoA from CO2 and at the same time makes ATP. This process is paid for by a proton gradient. This has been described by Shock as "a free lunch you get paid to eat". The energetics of the present acetyl CoA pathway based on the overall reaction below show an approximate DG0 value of -59 kJ/mol which can drive ATP synthesis.
2CO2 + 4H2 + CoASH --> CH3COSCoA + 3H2O.
The concentration of carbon dioxide in the primordial ocean was 1000 times higher than now. Vents produced large amounts of methane and hydrogen gas. There was little oxygen and hence lots of Fe2+. The enzymes involved in this acetyl-CoA pathway of carbon fixation have FeS clusters. It has also been shown that bubbles (which are really membrane bound spaces) of FeS and NiS can be made in deep sea vents. These could not only encapsulate precursor molecules but also serve as catalysts. Vents also can catalyze the fixation of nitrogen (to ammonia) and laboratory studied show that FeS can catalyze the conversion of formate (found in vents) into pyrimidines and purines. The studies of present methanogens and vent chemistry suggest that the critical ingredients and conditions for development of the first biological cells probably occurred in the vents.
To produce polymers, an energy source and monomers must exists. Concentration gradients found in simulations of vents produce million fold concentrated molecules. The transient heating and cooling of any double-stranded nucleic acids could lead to concentration amplification by a PCR like strand separation followed by reannealing. In addition, these vent regions possess a powerful, reoccurring energy source, a pH gradient, as the alkaline vented material entered the acidic oceans that exists with high CO2 concentrations, creating a gradient across an inorganic membrane. This is startlingly analogous to the pH gradient across membranes (acidic outside, alkaline inside) driven by the membrane complexes in the mitochondria and bacteria. Lane et al argue that the existence of membrane proton gradients as an energy source in all cells (eukaryotes, bacteria, and archaea) and in chloroplasts, mitochondria, corroborate their hypothesis. Bacteria and archaea share homologous ATPases and electron carriers (ferredoxins, quinones, and cytochromes). These similarities contrast to the differences in enzyme structures in fermentative pathways. Arguments that proton pumps evolved to pump proteins (and reduce pH gradients) can't explain their ubiquitous presence even in organisms not subjected to low pH. Hence the ubiquity of proton pumps supports the conjecture that they arose from the first protocells, possible comprised of inorganic walls and ultimately with amphiphilic molecules synthesized from precursors.
Creationists would argue that it would be impossible to evolve a structure with the complexity of membrane ATPase (which serve to collapse a pH gradient as the power the synthesis of molecules with large negative DG0 of hydrolysis). Lane et al propose that the earliest cells evolved ATPase like molecules in alkaline vents where pH gradients analogous to those in cells today arose. They envision cell-like columns lined by FeS membrane like structure with alkaline conditions inside and acid conditions outside. Nonpolar or amphiphilic molecule would line the inside of the cells/columns. A ATPase-like system could then take advantage of the pH gradient which constantly replenishes itself. If structures as complicated as ribosomes evolved from a subsequent RNA world, surely ATPase-like molecules could also. Other chemistry might have evolved earlier to utilize the energy source provided by the pH gradient.
If life originated in the vents, it would need an energy source to leave the vents. Presumably it would have evolved one to utilized pH gradient to replace the one it left in the alkaline vents. The substrate level phosphorylation of glycolysis that requires ATP input to make ATP would not provide the energy source needed. Cells that left would have had to produce their own proton gradient. Perhaps all the was needed initially was concerted conformational changes in proteins that upon exposure of a different pH changed their shape inducing pKa shifts in adjacent proton donors/acceptors leading to vectorial discharge of protons across a membrane. Perhaps the method described above in protocells was sufficient.
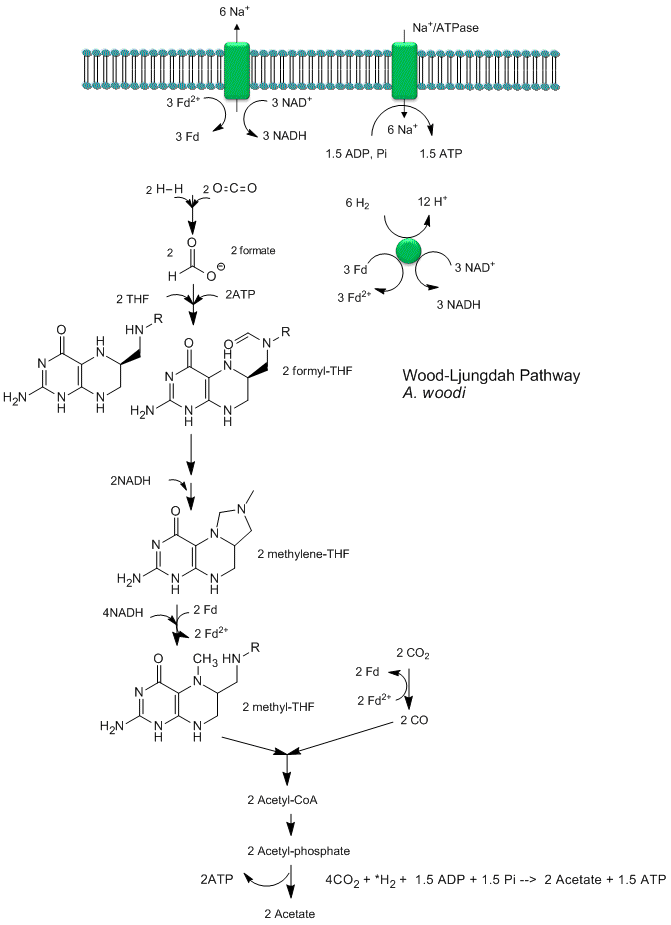
Recent analyses by Poehlein et a show that CO2 reduction (fixation) can be coupled to the production of a sodium ion gradient, which could collapse to drive ATP synthesis. Analysis of the genome of a gram positive bacteria, Acetobacterium woodii, an acetogen, shows the it has an ancient pathway for production of acetyl-CoA that can, in an anabolic fashion form biomass or in a catabolic fashion be cleaved to acetate with the production of ATP. It does not require classic electron carriers like ubiquinone or cytochrome C linked to protein gradient formation to drive ATP synthesis. Rather it has only a ferredoxin:NAD+ oxioreductase which couples oxidation to the formation of a sodium ion gradient, which collapses through an sodium ion transporter/ATP synthase to drive ATP synthase. A plausible reaction scheme based on genomic analysis is shown below:
Figure: Acetyl-CoA Synthase and Acetogenesis
Navigation
Return to Chapter 10: The Origin of Life Sections
Return to Biochemistry Online Table of Contents
Archived version of full Chapter 10: The Origin of Life

Biochemistry Online by Henry Jakubowski is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.