Biochemistry Online: An Approach Based on Chemical Logic

CHAPTER 2 - PROTEIN STRUCTURE
D: PROTEIN FOLDING AND STABILITY
BIOCHEMISTRY - DR. JAKUBOWSKI
Last Update: 3/1/16
|
Learning Goals/Objectives for Chapter 2D: After class and this reading, students will be able to
|
D1. INTRODUCTION
We've seen many static images of lipids and their aggregates as well as proteins. To think about how proteins fold, we have to think dynamically. Luckily we have the tools of molecular dynamics (MD) at our fingertips which helps us imagine how these processes take place and concomitantly how to probe protein folding experimentally. View the following two MD simulations and compare the spontaenously formation of a micelle and the folding of a protein before we delve into the complex topic of protein folding and stability.
Given the number of possibly nonnative states, it is amazing that proteins fold to the native state at all, let alone in a reasonable time frame. Consider this greatly simplified view of protein folding for a protein containing 100 amino acids. If each amino acid can adopt only 3 possible conformations, the total number of conformations could be 3100 = 5 x 1047. Assuming that it would take 10-13s to change each conformation, the time required to "test" all conformations would be 5 x 1034s or 1027 years, longer than the age of the universe (14 x 109 yr). Yet the protein can fold within seconds. This paradox is called the Levinthal paradox, after Cyrus Levinthal.
Lubert Stryer (in his classic Biochemistry text), shows a way out of this dilemma by using an analogy of a monkey sitting at a typewriter, and typing this line out of Hamlet: "Me thinks it is like a weasel." Random typing would produce that line after 1040 keystrokes on average, but if the correct letters were maintained, the number of keystrokes would be in the realm of a few thousand. Proteins could fold more quickly if they retain native-like intermediates along the way. Also remember that much of conformation space is already restricted by allowed phi/psi angles. (Remember the blank areas in the Ramachandran plot?)
Before we study the classic experiment of protein folding conducted by Anfinsen, study the simpler analogy below:
Figure: Socks and protein folding
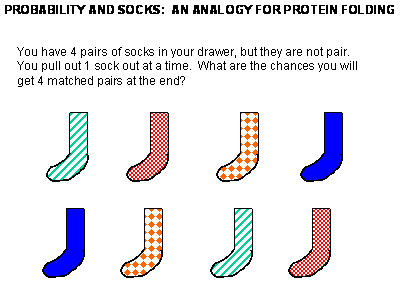
The classic experiment of Anfinsen has shown that, at least for some proteins, all the necessary and sufficient information required to direct the folding of a protein into the native state is present in the primary sequence of a protein. Anfinsen studied the in vitro (outside the cell, as opposed to in vivo, which is inside the cell, tissue, organ) folding of a single chain protein, RNase, which has 4 intrachain disulfide bonds.
Figure: RNase A with 4 Disulfide Bonds in red (image with VMD)
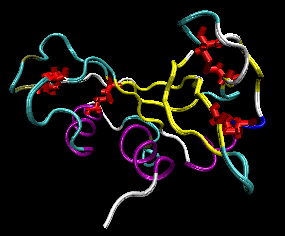
We have previously discussed how chemical agents (such as beta-mercaptoethanol, a disulfide reducing agent) can covalently interact with specific protein functional groups. Other substances can bind through complementary intermolecular forces to the active site or other cavities on the surface. Other reagents, like urea, acting through generalized solvent changes or nonspecific interactions with the protein, can alter protein folding. Anfinsen used two different reagents, 8 M urea and beta-mercaptoethanol, in combination to unfold, or denature, RNase to the nonnative or denatured state. He then removed the bME using dialysis, allowing the disulfides to reform. Next he removed the denaturing reagent, urea. To monitor if the protein was correctly refolded or renatured, he tested the activity of the protein compared to native protein. He found that the "refolded" protein retained only 1% of its initial activity. If, however, he added a catalytic amounts of bME, the protein soon retained 100% of its initial activity. For his work, he was awarded the Nobel Prize in Chemistry in 1972.
Figure: Anfinsen Experiment: Folding of RNase
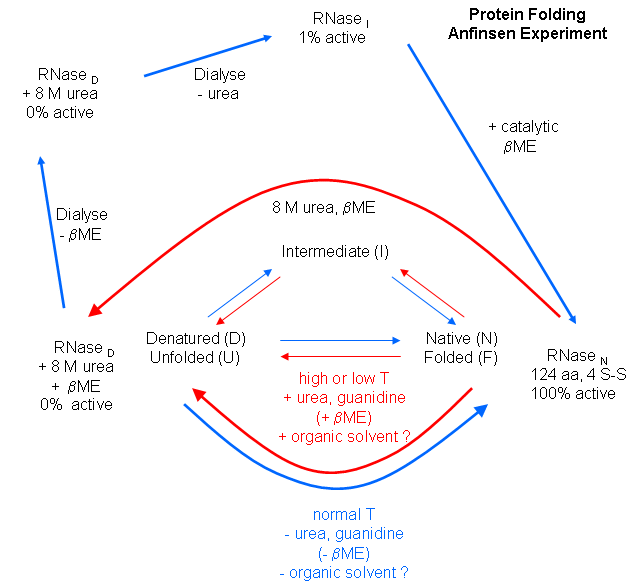
Figure: CATALYTIC SHUFFLING OF DISULFIDES WITH BETA-MERCAPTOETHANOL
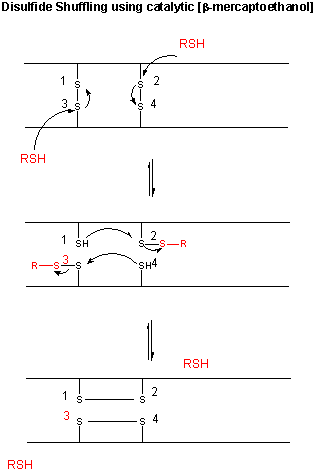
![]() Jmol: Updated
RNase A
Jmol14 (Java) |
JSMol (HTML5)
Jmol: Updated
RNase A
Jmol14 (Java) |
JSMol (HTML5)
Scientists have investigated the folding of proteins both in
vitro and in vivo. In vitro experiments involve denaturing the protein with
urea, guanidine hydrochloride, or heat, then refolding the protein by
removing the perturbant (denaturing agent), using spectral techniques to
follow the process. In vivo experiments involve the study of intracellular
proteins that assist folding. The in vitro experiments involve unfolding the
native state and then refolding it, while the in vivo ones involve folding
of the newly synthesized protein. An understanding of protein folding
can not be separated from an understanding of protein stability, and an
understanding of the nature of the native and denatured state.
In studying protein folding and stability/structure of the native and denatured states, both equilibrium (thermodynamic) and timed (kinetic) measurements are made. Folding occurs in the ms to second range, which limits the ability to study the presence of intermediates in the process. Some clever methods have been developed to study intermediates in protein folding by trapping specific intermediate structures, and investigating their structure and stability in a "leisurely" fashion. Alternatively, intermediates can be studied as they occur using stop flow kinetics. In this technique, a protein under denaturing conditions is rapidly mixed with a solution containing no denaturant or protein by injecting both solutions into a mixer/cuvette using syringes. The denaturant in the protein solution is now diluted such that renaturation can occur. Spectral measurements can begin at once.
A diagram summarizing these methods is shown below. Study it in conjunction with the text which follows.
Figure: Kinetic and thermodynamic measurements of proteins stability and folding
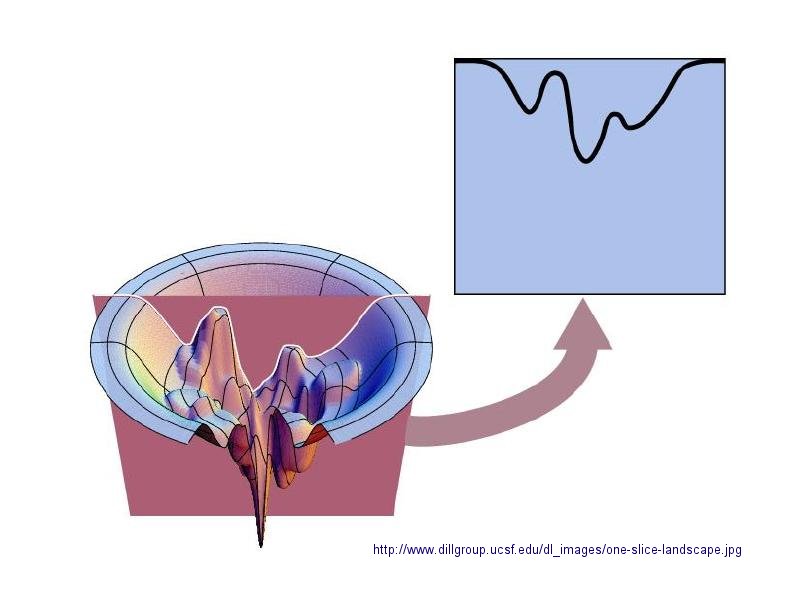
In considering the folding pathway, we will consider that the native protein represents the global energy minimum. All other states represent variations of the denatured state. Some, closer in energy to the native state, could be considered intermediates in the folding process. Instead of considering a folding "trajectory", consider protein folding occurring within a large folding landscape of free energy. Folding appears to proceed not by an obligatory pathway but a probabilistic or stochastic search of possible conformation. The free energy landscape must be shaped somewhat like a funnel such that a proteins could adopt a "reasonable" number of conformations which lead to the native state. Evolution has surely selected for sequences that can make it to that state. Localized secondary structure motifs (like a short alpha helix and beta turns) can form quickly (about 1 ms). Small proteins folding occurs, depending on the structure, over a wide time frame (ms to minutes). Mostly likely, a small number of amino acids coalesce into a core which nucleates folding into structures that are similar to the native state. Finally packing interactions collapse the structure into the native state.
In general, the more complex the fold of the backbone, the longer it takes the protein to fold. If complexity requires more interactions among distal regions of the polypeptide change, then the more complex the fold, the less probable that random interactions would lead to quick protein folding. The mechanisms of folding for larger proteins (greater than 100 amino acids) appear to proceed through intermediates, suggesting that different domains of the protein can fold independently.
Figure: Protein Folding Landscape: One View from Ken Dill
D2. Protein Folding In Vitro
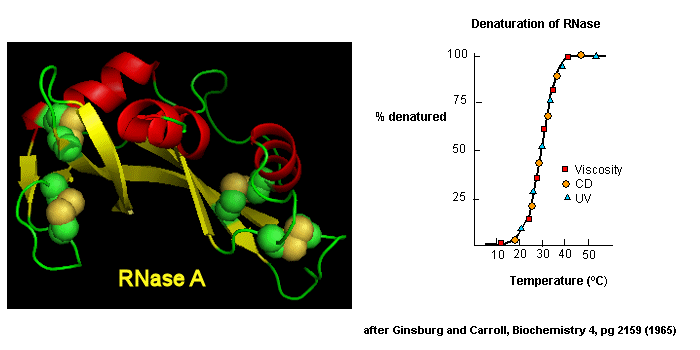
Early studies of protein folding involved small proteins which could be denatured and refolded in a reversible fashion. A two state model, D <===> N, was assumed. The denaturants were heat, urea, or guanidine HCl. Since the denatured states are less compact than the native state, the viscosity of the solution can be used as a measure of denaturation/renaturation. Likewise, the amino acid side chains in the differing states would be in different environments. The aromatic amino acid Trp, Phe, and Tyr absorb UV light. After excitation, the electrons decay to the ground state through several processes. Some vibrational relaxation occurs, bringing the electrons to lower vibrational energy levels. Some of the electrons can then fall to various vibrational levels at lower principle energy states through a radiative process. The photons emitted are lower in energy and hence longer in wavelength. The emitted light is termed fluorescence. The wavelength of maximum fluorescent intensity and the lifetime of the fluorescence decay is very sensitive to the environment of the amino acids. Hence fluorescence can also be used to measure changes in protein conformation. Other spectral techniques like CD spectroscopy as well as simple absorbance measurements, are used. For small, single domain proteins (such as RNase) undergoing reversible denatuation, graphs showing the extent of denaturation using each technique above, are superimposable, giving strong validity to the two state model.
Figure: Reversible denaturation
Proteins that fold without easily discernable, long lived intermediates and following a simple two state model, D <=> N are said to undergo cooperative folding. This simple model needed to be expanded as more proteins were studied. Some intermediates in the process were detected.
- Some proteins show two steps, one slow, one quick, in refolding studies, suggesting an intermediate. The longer a protein is kept in the denatured state, the more likely it is to display an intermediate. One accepted explanation for this phenomena is that during an extended time in the D state, some X-Pro bonds might isomerize from trans to the cis state, to form an intermediate. Alternatively, as in the case of RNase, which has a cis X-Pro bond in the native state, denaturation causes an isomerization to the trans state. In the case of RNase, to refold, the accumulating intermediate I must reisomerize in a slow step to the cis state, followed by a quick return to the N state.
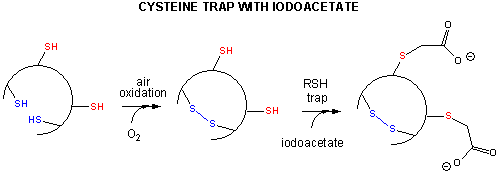
- Some proteins which contain multiple disulfide bonds that must reform correctly after reductive denaturation can refold into intermediates with the wrong S-S partner. Such intermediates can be trapped by stopping further S-S formation during refolding with the addition of iodoacetamide.
Figure: addition of iodoacetamide
As an example consider the following data on bovine pancreatic trypsin inhibitor.
Figure: Bovine pancreatic trypsin inhbitor (BPTI): Folding Kinetics - only native disulfide structures seem to form.
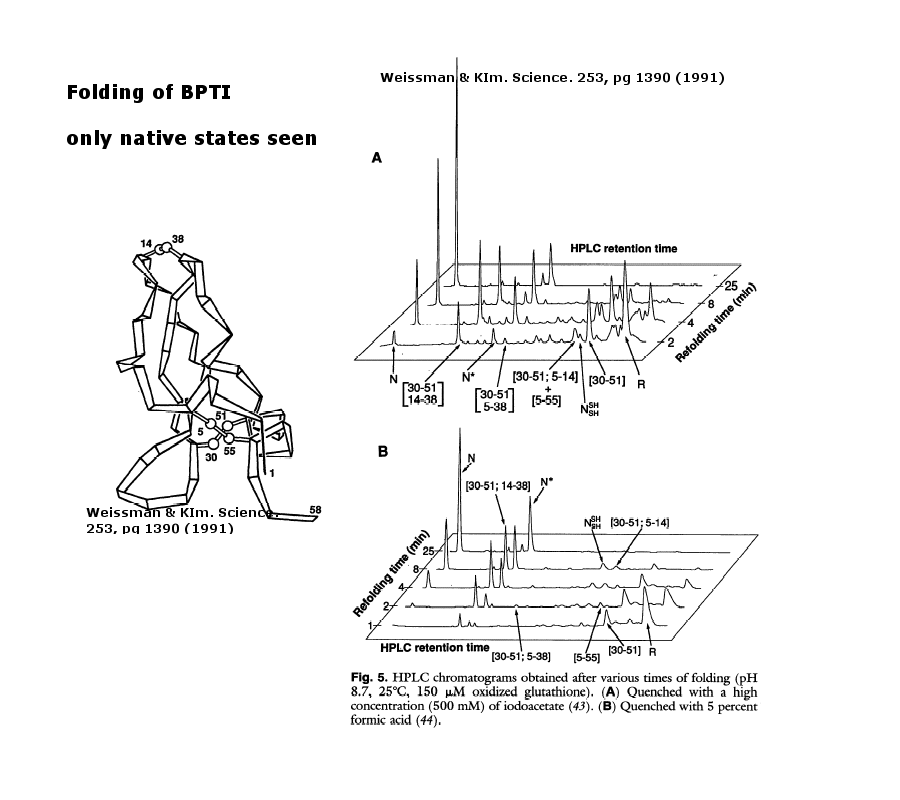
Figure: BPTI Folding Pathway In Vitro - gives possible scheme of folding intermediates
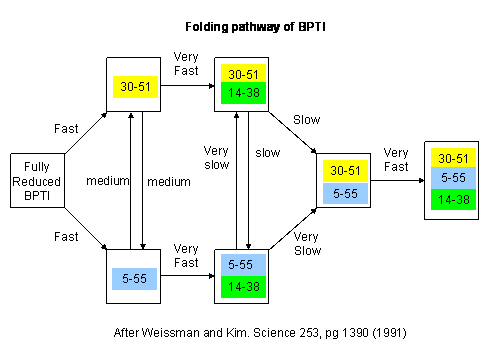
- Some proteins form partially folded but stable intermediates when folded under partially denaturing conditions. A good example is lactalbumin, which under mildly acidic conditions (pH 4), low levels of guanidine HCl, or neutral pH and low ionic strength in the absence of calcium (which normally binds to the protein), forms a stable, isolatable intermediate (I) called the molten globule (MG). The image below shows the folded state with two calcium ions bound.
Figure: lactalbumin (image made with Pymol)
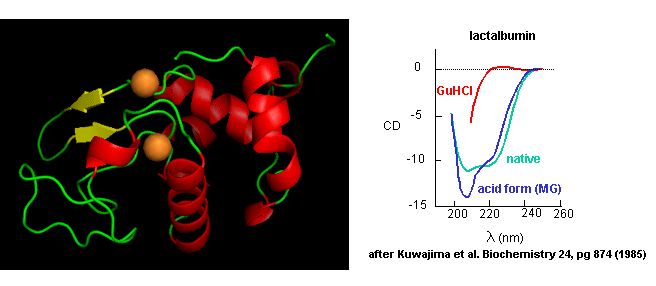
Data show that the MG is about 50% larger in volume than the N state. This compares to the denatured state, which can be 300% larger than the native state. Hence, it is more like the native state as studied by hydrodynamic techniques, but with more solvent accessibility of hydrophobic side chains. The MG has a similar CD spectra as the native state, but the aromatic side chains display the same UV absorption and fluorescent characteristics as the protein in 6 M guanidine HCl, suggesting that the final tertiary state has not yet completely formed. The secondary structure in the MG may not be the same as in the native state
NMR can also be used to detect folding intermediates. Using this technique, proteins are unfolded in D2O, which will cause the exchange of all Cs with ionizable protons, including, the amide Hs. An amine is a weak base (pKb around 3.5) so its conjugate acid, the protonated amine, has a pKa of around 9.5. An amide or peptide bond would be a weaker base than an amine since it's lone pair is less available (due to delocalization through resonance) for sharing with a proton. The pKa for the conjugate acid of the amide (in which the amide N is protonated and has a plus charge) is much lower, around -0.5, than the pKa for the conjugate acid of an amine. At 2 pH units greater than its pKa, the charged amide N is close to 100% deprotonated The pka of the protonated group is important since the rate of H exchange is related to the pKa, holding other variables constant. The pka of an unprotonated amine (RNH2 -> RNH- is very high (30s) and hence deprotonation of the RNH2 amine to form RNH- is not likely under normal conditions.
Figure: Exchange of all Cs with ionizable protons, including the amide Hs
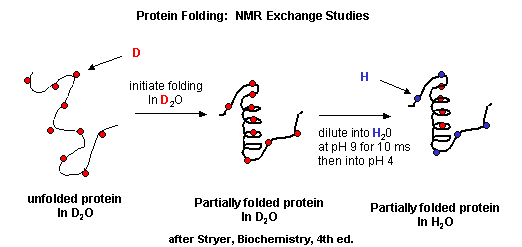
Refolding is initiated by diluting the protein into a solution without the denaturatant, but still in D2O. As the protein folds and becomes more compact, the buried atoms are now sequestered from the solvent, and no longer readily exchange Ds. Then the protein is placed in H2O at pH 9.0 for 10 ms, after which the pH is changed to pH 4.0. D --> H exchange is promoted at high pH, and quenched for the amide Ds and Hs at low pH. Amide H's that continue to exchange must be accessible to water. Those that aren't are usually buried in secondary structure.
Figure: Experimental data on model proteins. How would you interpret these graphs.
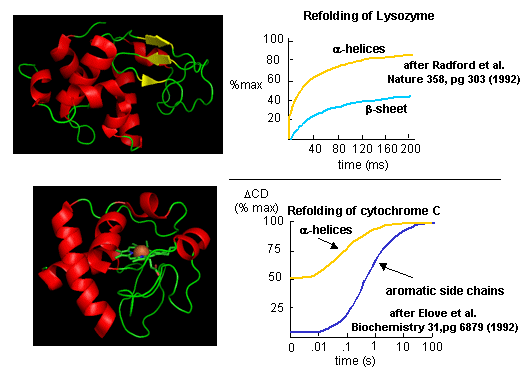
When the same techniques are applied to large, multidomain or oligomeric proteins, only a few percent refold in vitro. Incorrect intermolecular interactions and heterogeneous aggregation seems to be the main problems which prevent correct protein folding in vitro.
![]() Jmol: Updated
Apolactalbumin (w/o Ca2+)/Hololactalbumin
Jmol14 (Java) |
JSMol (HTML5)
Jmol: Updated
Apolactalbumin (w/o Ca2+)/Hololactalbumin
Jmol14 (Java) |
JSMol (HTML5)
D3. Folding of Single Protein Molecules
Protein folding/unfolding reactions can now be studied on single protein molecule using an optical tweezer, as illustrated in the figure below based on a study of RNase H from E. Coli (Cecconi et al, 2005). RNaseH was mutated to contain two Cys residues at positions 4 and 155 (near the ends of the chain). This allowed the protein to be covalently linked through the free sulfhydryl of Cys to two dsDNA molecules (500 bp) chemically modified at one end to form covalent bond to the Cys- containing protein. The CD spectra of the protein linked to the DNA tether was identical to the unmodified protein. The DNA linkers were linked to two different styrene beads though different chemistries to produce the structure shown below.
Figure: Unfolding of RNase H with Optical Tweezers
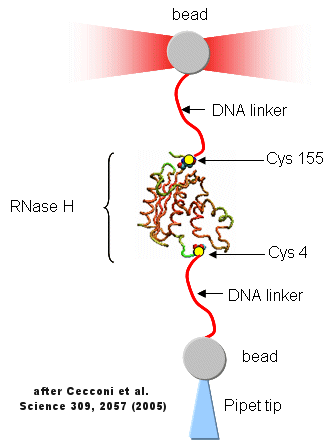
The enzyme was active as shown by enzyme activity measurements. The protein can be stretched and contracted by moving the bead connected to the pipet relative to the bead in the optical trap. As the force (measured in piconewtons) is increased, the molecule is stretched. As a control, the two beads were connected directly with DNA handles without the protein. In the control case, a graph showing extension between the beads vs force increases slowly in a curvilinear fashion at first and then linearly. When the protein is inserted, differences are observed. Upon stretching and relaxing two transitions (shifts) were observed with RNase H that are not seen with DNA alone. These occurred at about 19 pN (an abrupt shift) and a smaller one at 5 pN. These transitions are consistent with the N→D and an I --> D transition , as shown in the figure below (adapted from Cecconi et al). These data are consistent with previous studies of this protein which show the the central, stable core of the protein folds first.
Figure: RNaseH Folding Transitions: Optical Trap
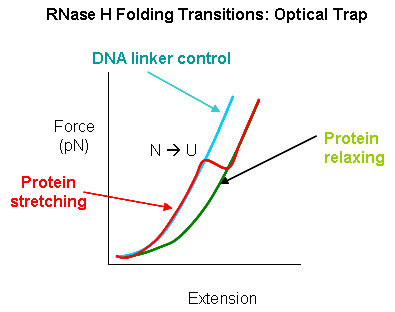
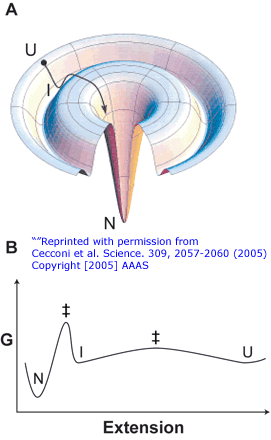
Figure: Folding Lanscape of RNase H: Optical Tweezers
D4. The Denatured State
Although the structure of native and native-like states can be determined using x-ray crystallography and in solution using NMR, little detailed information exists on the actual structure of denatured and intermediate states. Intermediate states are difficult to trap in a way that allow details structural analysis. In contrast to the "native" state which consists of an ensemble of closely related states, intermediates and denatured states would consist of an ensemble of many different states, making structural analysis more difficult. Religa and others from Fersht's lab have engineered a mutant of the engrailed homeodomain (En-HD) from Drosophila melanogaster that allows such structural analyses to be performed. The mutation, Leu16Ala (L16A), destabilizes the protein such that it can be denatured simply by changing ionic strength. It is stable at high ionic strength and folds quickly under those condition. However at physiological ionic strength it is "denatured" but contains significant alpha-helical structure but has nonnative contacts. It behaves like an early folding intermediate in that if placed in solutions of higher ionic strength it rearranges to form the native state. If placed in lower ionic strength, it progressively "unfolds" to yet other states. Given the ambiguities in how to define denatured and early folding intermediates states, Ferscht's group suggest an "explicit" definition of the denatured state. They define the unfolded state (U) as the "maximally unfolded state of a protein, in which the backbone NH groups have little protection against 1H/2H exchange". They define the denatured state, D, as the "lowest energy non-native state under a defined set of conditions". In this scenario, the denatured state could also be a folding intermediate if placed in conditions that promote folding. Previous work from the group showed that the denatured state of En-HD has three helices protected from 2H exchange and was one kcal/mol lower in energy than the unfolded state.
![]() Jmol:
Updated Engrailed homeodomain - denatured and native state
Jmol14 (Java) |
JSMol (HTML5)
Jmol:
Updated Engrailed homeodomain - denatured and native state
Jmol14 (Java) |
JSMol (HTML5)
D5. Multiple Conformations from Same Sequence
1. Silent Single nucleotide polymorphisms (SNPs): For some amino acids, multiple triplet nucleotide sequences (codons) in the coding regions of a gene for a protein lead to the incorporation of the same amino acid in the protein sequence. Hence two proteins identical in amino acid sequence might have slightly different nucleotide sequences in the gene that encodes them. Such single nucleotide polymorphisms (SNPs) in coding regions were thought to have no effect on the tertiary structure and biological function of a protein if the single nucleotide variation did not lead to the insertion of a different amino acid into the growing peptide chain (i.e the codons were synonymous and the mutations presumably silent with no effect). Recently single nucleotide polymorphisms (SNPs) in the gene for the product of the MDR1 (multidrug resistance 1) gene, P-glycoprotein, was shown to result in a protein with different substrate specificity and inhibitor interactions, and hence a different 3D structure. One possible explanation for this observation is a difference in the rate of translation of the mRNA for this membrane protein. Different rates might lead to different intra- and intermolecular associations, which could lead to different final 3D structures as the protein cotranslationally folds and inserts into the membrane. This would especially be true if two possible structures where close enough in free energy but separated by a significant activation energy barrier, precluding simple conformational rearrangement of one conformation to another.
2. Metamorphic Proteins: In addition to prion proteins, it appears that many proteins can adopt more than one conformation under the same set of conditions. In contrast to prion proteins, however, in which the formation of the beta-structure variant is irreversible since the conformational change is associated with aggregation, many proteins can change conformations reversibly. Often, these changes do not appear to be associated only with binding interactions that trigger the change. Murzin has described proteins that change conformations on change of pH (viral glycoproteins), redox state (chloride channel), disulfide isomerization (lysozyme), and bound ligand (RNA polymerase as it initiates and then elongates the growing RNA polymer). He cites two proteins that appear to changes state without external signals. These include Mad2, in which the two conformers share extensive similarity, and Ltn10 (lymphotactin), in which they don't. One form of lymphotactin (Ltn 10) binds to similar lymphokine receptors, while the other (Ltn 40) binds to heparin. Folding kinetics may play a part in these examples as well, as proteins capable of folding to two conformers independently and quickly might prevent misfolding and aggregation that might occur if they had to completely unfold first before a conformational transition. Both Mad2 and Ltn10 alter conformation through transient formations of dimers, which facilitate conformational changes without widespread unfolding. Mutations in Ltn10 can cause the protein to adopt the Ltn40 conformation, Hence primordial "metamorphic" proteins could, by simple mutation, produce new protein functionalities.
3. Intrinsically Disordered Proteins (IDPs): Many examples of proteins that are partially or completely disordered but still retain biological function have been found. At first glance this might appear to be unexpected, since how could such a protein bind its natural ligand with specificity and selectivity to express its function? Of course one could postulate ligand binding would induce conformational changes necessary for function (such as catalysis) in an extreme example of an induced fit of a ligand compared to a "lock-and-key" fit. Decades ago, Linus Pauling predicted that antibodies, proteins that recognize foreign molecules (antigens), would bind loosely to the antigen, followed by a conformational change to form a more complementary and tighter fit. This was the easiest way to allow for a finite number of possible protein antibodies to bind a seemingly endless number of possible foreign molecules. This is indeed one method in which antibodies can recognize foreign antigens. Antibodies that bind to antigen with high affinity and hence high specificity more likely bind through a lock and key fit. (Pauling, however, didn't know that the genes that encode the proteins chains in antibodies are differentially spliced and subjected to enhanced mutational rates which allow the generation of incredible antibody diversity from a limited set of genes.)
It's been estimated that over half of all native proteins have regions (greater than 30 amino acids) that are disordered, and upwards of 20% of proteins are completely disordered. Regions of disorder are enriched in polar and charged side chains which follows since these might expected to assume many available conformations in aqueous solutions compared to sequences enriched in hydrophobic side chains, which would probably collapse into a compact core stabilized by the hydrophobic effect. Mutations in the disordered regions tend to preserve the disordered region, suggesting that the disordered region is advantageous for "future" function. In addition, mutations that cause a noncoding sequence to produce a coding one invariably produce disordered protein sequences. Disordered proteins tend to have regulatory properties and bind multiple ligands, in comparison to ordered one, which are involved in highly specific ligand binding necessary for catalysis and transport. The intracellular concentration of disordered proteins has also been shown to be lower than ordered proteins, possibly to prevent occurrences of inappropriate binding interactions mediated through hydrophobic interactions, for example. Processes to accomplish this include more rapid mRNA and protein degradation and slower translation of mRNA for disordered proteins. For a similar reason, misfolded proteins are targeted for degradation as well. Figure A below shows the mean net charge vs the mean hydrophobicity for 275 folded and 91 natively unfolded proteins. Figure B shows the relative amino acid composition of globular (ordered) proteins compared to regions of disorder greater than 10 amino acids in disordered proteins. The two different grey bars were obtained with two different versions of the software used to analyze the proteins. Again the graph shows an enrichment of hydrophilic amino acids in disordered proteins.
Figure: Characteristics of Intrinsically Disordered Proteins
from open access journal: Dunker, A. et al. BMC Genomics 2008, 9(Suppl 2):S1 doi:10.1186/1471-2164-9-S2-S1
Many experimental methods can be used to detect disordered regions in proteins. Such regions are not resolved well in X-Ray crystal structures (have high B factors). NMR solution structures would show multiple, and differing conformations. CD spectroscopy likewise would show ill-defined secondary structure. In addition solution measurements of size (light scattering, centrifugation) would show larger size distributions for a given protein.
What types of proteins contain disorder? The above experimental and new computational methods have been developed to classify proteins as to their degree of disorder. There appears to be more IDPs in eukaryotes than in archea and prokaryotes. Many IDPS are involved in cell signaling processes (when external molecules signal cells to respond by proliferating, differentiating, dying, etc). Most appear to reside in the nucleus. The largest percentage of known IDPs bind to other proteins and also to DNA. These results suggest that IEPs are essential to protein function and probably confer significant advantages to eukaryotic cells as multiple functions can be elicited from the interaction of a single IEP (derived from a single gene) with different protein binding partners. This would greatly extend the effective genome size in humans, for examples, from around 25,000 with specified function, to many more. This doesn't even take into account the increase functionalities derived from post-translational chemical modifications.
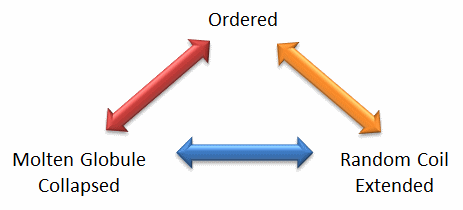
We will discuss intrinsically disordered proteins further in Chapter 5. What is clear from recent finding is that protein structure is fluid and complex and our simple notions and words to denote proteins as either native or denatured are misguided and constrain our ideas about how protein structure elicits biological function. For example, what does the word "native" mean, if proteins exist in multiple states in vivo and in vitro simultaneously? Dunker et al (2001) have coined the concept "Protein Trinity" to move past the notion that a single protein folds to a single state which elicits a single function. Rather each of the states in the "trinity", the ordered, collapsed (or molten globule) and extended (random coil) coexist in the cell. Hence all can be considered "native" and all contribute to the function of the cell. A single IDP could bind to many different protein partners, each producing different final structures and functions. IDPs would also be more accessible and hence susceptible to proteolysis, which would lead to a simple mechanism to control their concentrations, an important way to regulate their biological activity. Their propensity to post-translational chemical modification would likewise lead to new types of biological regulation.
Figure: The Protein Trinity: Ordered, Collapsed and Extended States
These ideas have profound ramifications for our understanding of the expression of cellular phenotype. In addition, a whole new world of drug target is available by finding drugs that modulate the transitions between ordered, collapsed and extended protein states. Likewise, side effects of drugs might be understood by investigating drug effects of these transitions in IDPs not initially targeted.
-
 PONDR
- Predictor of Naturally Occurring Disorder
PONDR
- Predictor of Naturally Occurring Disorder -
 Database
of Protein Disorder
Database
of Protein Disorder
4. Catalysis by Molten Globule: A recent example (Bemporad) that a bacterial acylphosphatase has catalytic activity as a molten globule further questions our notions of structure and enzyme activity. In this example, substrate interaction did not induce global conformational changes in the protein. Molecular dynamics simulations showed that many partially disordered conformations of the protein are present, and the disorder involved the active site. However, parts of the protein are more ordered and form a "scaffold" which keeps the catalytic and substrate binding amino acids near enough that binding could engender conformational rearrangements at the active side and subsequent catalytic activity.
D6. Protein Folding In Vivo
There are many differences between how a protein might fold or unfold in a cell compared to a test tube.
- The total concentration of all the proteins and nucleic acids in cells are estimated to be about 350 g/L, or 350 mg/ml. Most measurement in the lab are conducted in the range of 0.1 to 10 mg/ml
- Proteins are synthesized in cells from an N to C terminal direction. Hence the nascent protein, as it emerges from its site of synthesis (the ribosome), might fold into intermediate structures since not all of the protein sequence is yet available to direct folding.
- Proteins are synthesized in the cytoplasm, but they have to find their final place in the cell. Some end up in membranes, some must translocate across one or even two different membranes to end up in specific organelles like the Golgi, mitochondria, chloroplasts (in plant cells), nuclei, lysosomes, peroxisomes, etc. Do they translocate in their native state?
Additional evidence suggests that protein folding/translocation requires assistance (i.e. catalysis) in the cell.
- Mutant cells defective in certain proteins can lead to the accumulation in the cells of misfolded and aggregated proteins.
- eukaryotic genes (taken from higher cells which contain nuclei and internal organelles), when transferred into prokaryotes (bacteria, like E. Coli), can be expressed to form protein, but they often misfold and aggregate in the bacterial cells and form structures called inclusion bodies.
Hence recombinant proteins expressed in vivo have the same problems in folding as larger proteins in vitro. In both cases, conditions favor accumulation of nonnative proteins with exposed hydrophobic groups leading to aggregation. Aggregation also occurs in vivo when a protein is over-expressed or expressed at a higher temperature than normal. Why? Mutant cells have been selected that actually suppress inclusion bodies in vivo. This effect was mediated by a class of proteins which are expressed by the bacteria and other cells when their temperature is raised. The function of these proteins, called heat shock proteins (Hsp), was unknown until it was realized that they facilitate correct protein folding, in part, by binding to denatured proteins in the cells before they aggregate into inclusion bodies. Further studies discovered a large number of proteins that seem to facilitate protein folding and prevent aggregation in vivo. These proteins are now called molecularchaperones. They are classified on the basis of their molecular weight) and can be divided into at least two families, the Hsp-70 chaperone family and the chaperonin and Hsp 90 families as illustrated and summarized in the figure and text below.
Hsp-70 Family: This family includes DnaK/DnaJ and GrpE proteins in prokaryotes and immunolglobulin heavy chain binding protein (BiP) and alpha crystalline in eukaryotes. Alpha crystalline comprises 30% of the lens proteins in the eye, where it functions, in part, to prevent nonspecific, irreversible aggregates. These proteins have molecular weights of about 70K and :
- bind to growing polypeptide chains as they are synthesized on ribosomes.
- express activity as monomers.
- have ATPase activity - i.e. they cleave the phosphoanhydride ATP (which can drive reactions).
- bind short, extended peptides, which stimulates the ATPase activity
- release bound peptides after ATP cleavage
A figure showing
In prokaryotes, a protein called trigger factor (TF) binds in a co-translational process to proteins as they begin to emerge from the ribosome and catalyzes correct folding of about 70% of bacterial protein. The rest requires additional chaperones, including DnaJ and DnaK which bind proteins during synthesis in a cotranslational process. Upon interaction with the DnaJ-bound protein, DnaK hydrolyzes bound ATP, resulting in the formation of a stable complex between DnaJ and DnaK. GrpE, a nucleotide exchange factor for DnaK, facilitates the releases ADP from DnaK. Rebinding of ATP to DnaK then triggers the release of the substrate protein. This cycle repeats itself until the protein is fully folded. For about 20% of proteins in E. Coli, the DnaK/DnaJ/GrpE cycle leads to complete post-translational folding of proteins . Eukaryotes utilize an analogous set of proteins Hsp70 complex proteins including If folding is still incomplete after several rounds, the fully synthesized yet incompletely folded protein interacts with an amazing catalyst of protein folding, the chaperonin system.
Chaperonins- including chaperonin 60 (or GroEL in E. Coli) and chaperonin 10 (or GroES in E. Coli) in chloroplasts, mitochondria and bacteria, and TCP-1 in eukaryotic cytoplasm.
These proteins:
- bind to proteins after they have left the ribosome or have been transported into organelles like mitochondria.
- express activity as multimers. GroEL consist of two stacks of rings of monomers, with 7 monomers in each ring (each monomer around 60K MW), forming a hollow cylinder. GroES consist of one single ring of 7 monomers (each 10K MW). The GroES complex forms a lid over one open end of the GroEL cylinder. Proteins can fold within the cavity in GroEL (lined with hydrophobic patches) without "fear" of aggregation. GroEL also binds and cleaves ATP, leading to conformational changes inside the barrel and hiding of the hydrophobic patches in Gro EL, which leads to the releases of the unfolded peptide. The process proceeds until the folding protein passes through the barrel and is released in its correct folded state. Do you find this amazing?
- bind nonnative proteins at the GroEL opening of a complex of GroEL and GroES, which has a large hydrophobic cavity.
- Molecular Chaperonins in Disease
GroEL has also been shown to bind in its hydrophobic cavity a fluorescent CdS semiconductor nanoparticle which can be released on addition and cleavage of ATP. There are two classes of chaperonins:
- Class I: Those found in bacteria, chloroplasts and mitochondria. The have structures analogous to GroEL (two rings of 7 identical monomers) and Gro ES.
- Class II: Those found in archebacteria and in the cytoplasm of eukaryotic cells. These contain two rings of 8-9 subunits which may not be identical.
![]() Jmol:
Updated GroEL/ES
Jmol14 (Java) |
JSMol (HTML5)
Jmol:
Updated GroEL/ES
Jmol14 (Java) |
JSMol (HTML5)
Other chaperons have proven to be of clinical significance. Hsp 90 is a chaperone that is expressed both in normal and tumor cells. It appears to have special importance in tumor cells in helping key proteins involved in malignancy (signal transduction proteins such as HER-2/ErbB2, Akt, Raf-1, Bcr-Abl, and p53) to maintain their shapes under conditions of drug exposure and the inherent genetic instability present in the cells. Drugs that bind to and inhibit Hsp90 appear to have much greater effect on tumor cells, making this protein a new chemotherapeutic target to treat cancer. Recent studies by Kamal et al. have shown the drug 17-AAG binds Hsp90 about 100 times as strongly in tumor cells than in normal cells. Hsp 90 appears to be complexed to other "co-chaperones" in the tumor cells which lead to higher drug binding affinity. The chaperone complex may actually induce the drug to adopt a different conformation. A comparison of chaperone catalyzed folding in prokaryotes and eukaroytes is shown below.
Figure: Comparison of cytosolic prokaryotic and eukaryotic chaperone pathways
Additional Proteins Which Catalyze Protein Folding: Chaperons function to minimize protein aggregation, which increases the efficiency of the entire process. Other proteins in the cell actually catalyze specific steps. Here are two examples:
- Protein Disulfide Isomerase (PDI) - catalyzes the conversion of incorrect to correct disulfides. The active site consists of 2 sets of the the following sequence - Cys-Gly-His-Cys, in which the pKa of the Cys are much lower (7.3) than normal (8.5). How would this facilitate disulfide isomerization?
- Peptidyl Prolyl-Isomerase (PPI) - catalyses X-Pro isomerization, by a mechanism which probably involves bending the X-Pro peptide bond. How would this facilitate the process?
Many proteins have been found to possess PPI activity. One class is the immunophilins. These are small proteins found in the cytoplasm that bind anti-rejections drugs used to prevent tissue rejection after transplanation. The immunophilin FK506 binding protein (FKBP) binds FK506 while the protein cyclophilin binds that anti-rejection drug cyclosporin. The complex of cyclophilin:cyclosporin or FKBP:FK506 binds to an inhibits calcineurin, an important protein (with phosphatase activity) in immune cells (T cells) required for T cell function. In this case, immunophilin:drug binding to calcineurin inhibits the activity of the T cell, preventing immune attack on the transplanted tissue, preventing rejection. The immunosuppressant drugs (FK506 and cyclosporin) inhibit the PPI activity of their respective immunnophilin. The extent to which the PPI activity of cyclophiin is required for its activity is unclear, but it seems to be important for some of its biological effects.
![]() Jmol:
Updated Cyclophilin
Jmol14 (Java) |
JSMol (HTML5)
Jmol:
Updated Cyclophilin
Jmol14 (Java) |
JSMol (HTML5)
- Animation: Folding and Degradation of Proteins in vivo
As the site responsible for folding of membrane proteins and proteins destined for secretion, as well as the major site for lipid synthesis, the endoplasmic reticulum (ER) must be able to maintain homeostatic conditions to ensure proper protein formation. Plasma cells that synthesize antibodies for secretion as part of the immune activation, show large increases in protein chaperones and ER membrane size
The main pathway controlling ER biology is the unfolded protein response (UPR) signaling pathway. If demand for protein synthesis in the ER exceeds capacity, unfolded proteins accumulate. This ER stress conditions activates a protein called IRE1, a transmembrane Ser/Thr protein kinases (which phosphorylates proteins). IRE1 activates a transcription factor that controls transcription of many genes associated with protein folding in the ER. Another protein, ERAD (ER-associated degradation) which moves unfolded proteins back into the cytoplasm where they are degraded by the proteasome. Proteins involved in lipid synthesis are also activated as lipids are needed for membranes as the ER increases in size. If the stress can not be mitigated the signaling pathway leads to programmed cell death (apoptosis).
Schuck at al investigated the specific role and importance of UPR in the homeostasis of ER as modeled by the yeast Saccharomyces cervisiae. The UPR signaling pathway was analyzed using light and electron microscopy to visualize and quantify ER growth under various stress conditions. Western blotting procedures were performed to determine chaperone protein concentrations after stress induction and association with ER expansion after the ER was exposed to various treatment conditions. The authors found ER membrane expansion occurred through lipid synthesis since stress induction increased concentrations of proteins responsible for promoting lipid synthesis and expansion failed when the proteins were absent and lipid concentration was low. In addition, these lipid synthesis proteins were activated by the UPR signaling pathway. By separating ER size control and UPR signaling, they found that expansion occurred regardless of chaperone protein concentrations. However, if lipid synthesis genes were not available, raising the ER chaperone level helped alleviate stress levels in ER.
D7. Redox Chemistry and Protein Folding
In general we envision the interior of a cell to be in a reducing environment. Cells have sufficient concentrations of "b-mercaptoethanol"-like molecules (used to reduce disulfide bonds in proteins in vitro) such as glutathione (g-Glu-Cys-Gly) and reduced thioredoxin (with an active site Cys) to prevent disulfide bond formation in cytoplasmic proteins. For disulfide bonds to occur in a protein, a free sulfhydryl reacts with another one on a protein to form the more oxidized disulfide bond. This reaction occurs more readily if one of the Cys side chains had a lowered pKa (due to its immediate environment) making it a better nucleophile in the reaction. Most cytoplasmic proteins contain Cys with side chain pKa > 8, which would minimize disulfide bond formation as the Cys are predominantly protonated at that pH.
Disulfide bonds in proteins are typically found in extracellular proteins, where they serve to keep multisubunit proteins together as they become diluted in the extracellular milieu. These proteins destined for secretion are cotranslationally inserted into the endoplasmic reticulum (see below) which presents an oxidizing environment to the folding protein and where sugars are covalently attached to the folding protein and disulfide bonds are formed (see Chapter 3D: Glycoproteins - Biosynthesis and Function). Protein enzymes involved in disulfide bond formation contain free Cys which form mixed disulfides with their target substrate proteins. The enzymes (thiol-disulfide oxidoreductases, protein disulfide isomerases) have a Cys-XY-Cys motif and can promote disulfide bond formation or their reduction to free sulfhydryls. They are especially redox sensitive since their Cys side chains must cycle between and free disulfide forms.
Intracellular disulfide bonds are found in protein in the periplasm of prokaryotes and in the endoplasmic reticulum (ER) and mitochondrial intermembrane space (IMS) of eukaryotes. For these proteins, the beginning stage of protein synthesis (in the cytoplasm) is separated temporally and spatially from the site of disulfide bond formation and final folding. Disulfide bonds can be generated in a target protein by concomitant reduction of a disulfide in a protein catalyst, leaving the net number of disulfides constant (unless the enzyme is reoxidized by an independent process). Alternatively, a disulfide can be formed by transfer of electrons to oxidizing agents such as dioxygen.
In the ER, disulfide bond formation is catalyzed by proteins in the disulfide isomerase family (PDI). To function as catalysts in this process, the PDIs must be in an oxidized state capable of accepting electrons from the protein target for disulfide bond formation. A flavoprotein, Ero1, recycles PDI back to an oxidized state, and the reduced Ero1 is regenerated on passing electrons to dioxygen to form hydrogen peroxide. In summary, on formation of disulfides in the ER, electrons flow from the nascent protein to PDIs to the flavin protein Ero1 to dioxgen (i.e. to better and better electron acceptors). The first step is really a disulfide shuffle, which, when coupled to the subsequent steps, leads to de novo disulfide bond formation.
In the mitochondria, disulfide bond formation occurs in the intermembrane space (IMS) and is guided by the �mitochondria disulfide relay system.� This system requires two important proteins: Mia40 and Erv1. Mia40 contains a redox active disulfide bond cys-pro-cys and oxidizes cys residues in polypeptide chains. Erv1 can then reoxidize Mia40 which can in turn get reoxized itself by the heme in cytochrome c. Reduced cytochrome C is oxidized by cytochrome C oxidase of electron transport through passage of electrons to dioxygen to form water. The importance of IMS protein oxidation is less understood, but it is believe that the oxidative stress caused by a dysfunction could lead to neurodegenerative diseases.
A recent review by Riemer et al compares the ER and mitochondrial processes for disulfide bond formation:
-
Many more and diverse proteins form disulfides in the ER compared to the IMS. Most in the IMS have low molecular mass and have two disulfide bonds between helix-turn-helix motifs. These protein substrates include chaperones that facilitate localization of proteins in the inner membrane, and in proteins involved in electron transport in the inner membrane.
-
There are many PDIs in the ER, probably reflecting the structural diversity of protein substrates in the ER. However Mia40 appears to be the only PDI in the IMS.
-
"De novo" disulfide bond formation is initiated by Ero1 in the ER and Erv1 in the IMS. Convergent evolution led to the similar structures for both - a 4-helix bundle that binds FAD with two proximal Cys.
-
The mitochondria pathway lead to water formation on reduction of dioxygen, not hydrogen peroxide, minimizing the formation of reactive oxygen species in the mitochondria. The peroxide formed in the ER is presumably convert to an inert form.
-
The IMS is in more intimate contact with the cytoplasm through outer membrane proteins called porins which would allow some glutathione access. The IMS presents a more oxidizing environment than the cytoplasm (with more glutathione). The ER, without a porin analog, would be more oxidizing.
-
Reversible formation of disulfides in the ER regulates protein activity.
Disulfide bond regulation in the Periplasmic Space of Bacteria
The redox sensitivity of the Cys side chain found in disulfide bonds is important in regulating protein activity. In particular, the thiol group of the amino acid Cys, an important nucleophile often found in active site, can be modified to control protein activity. The formation of a disulfide bond or the oxidation of free thiols to sulfenic acid or further to sulfinic or sulfonic acid can block protein activity. The E. Coli periplasmic proteins DsbA (disulfide bond A) converts adjacent free thiols into disulfide-linked Cystine, in the process becoming reduced. DsbB reoxidized DsbA back to its catalytyically active form. What about periplasmic protein like YbiS with an active site Cys? Since the environment of the periplasm is oxidizing, YbiS mist be protected from oxidative conversion of the free Cys to either sulfinic or sulfonic acids causing the protein to become inactive. The mechanism involves two periplasmic proteins known as DsbG and DsbC which are similar to thioredoxin. These two proteins are able to donate electrons to the unprotected thiol preventing it from becoming oxidized, which allows YbiS to remain active in the periplasm. To maintain activity, DsbG and DsbC are reduced by another periplasmic protein, DsbD.
D8. Protein Transport Across Membranes
How does a protein "decide" its final location after synthesis? Protein synthesis occurs in the cytoplasm, but proteins may end up outside of the cell, in cell membranes, internalized into various organelles, or remain in the cytoplasm. How is the decision made? There must be signals in the protein which target proteins to various sites in a cell, where processing can occur. Proteins that are destined for secretion or plasma membrane insertion typically have a signal peptide at the N-terminus which binds to a signal recognition particle in a cotranslational process which temporarily arrests translation. This complex docks to signal recognition complex docking sites in the endoplasmic reticulum membrane, where translation continues as the nascent polypeptide extends through a protein pore in the ER membrane. Gunter Blobel won the Noble Prize in Medicine in 1999 for "for the discovery that proteins have intrinsic signals that govern their transport and localization in the cell".
Figure: Overview - Synthesis of Protein Destined for Secretion
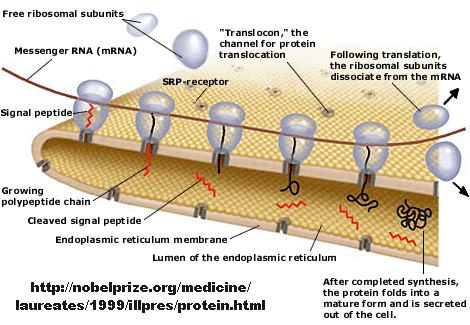
Figure: Signal Recognition Particle Complex
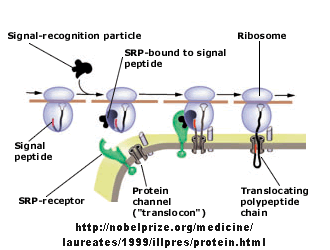
Figure: Sec Dependent Pathway for Post-translational translocation
(reprinted with permission from Kanehisa Laboratories and the KEGG project: www.kegg.org )
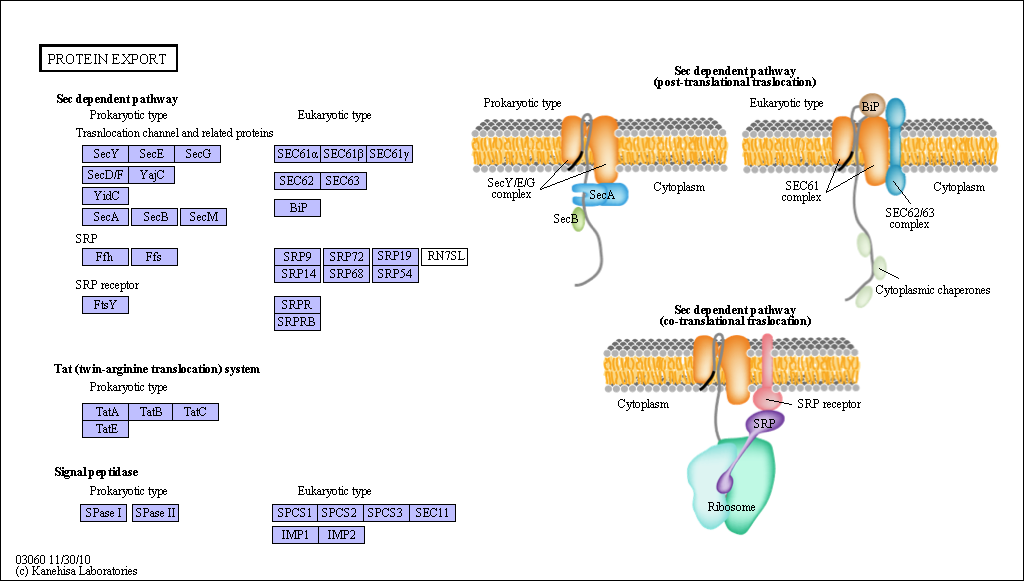
If destined for secretion, it enters the lumen of the ER. Proteins destined for insertion into the cell surface membrane gets "stuck" in the ER membrane, and through a process of vesiculation merges with the Golgi and eventually with the cell surface membrane. Proteins that are taken into organelles like mitochondria are done so in a post-translational process that requires facilitation by protein chaperones. Final protein folding occurs inside the organelle. In both cases, nonnative proteins pass through the membrane after which final folding occurs.
An intriguing question is how the decision is made to keep a protein either in the membrane or allow it to pass through completely (in the case of proteins destined for secretion). Hessa et al investigated this "decision-making" process by studying the eukaryotic membrane pore protein complex, Sec 61 translocon (show in the above figures), whose activity must be closely regulated with the folding of the growing protein. In studying this process, they considered three local regions in a membrane: the hydrophobic region comprised of the nonpolar acyl tails of membrane lipids, the interfacial region in the vicinity of the polar head groups, and the aqueous regions (bulk water) on each side of the head groups. A 19 amino acid peptide was used as the experimental model protein which was added to the translocon. This size was chosen since it is just long enough to span the hydrophobic part of the membrane if the peptide were in an alpha-helical conformation (which is common in membrane-spanning proteins). They varied the proportion of amino acids that tend to partition into each of three regions and studied the disposition of the peptide after interaction with membrane and translocon. To test if the results were consistent with the thermodynamics of amino acid partitioning into nonpolar environments (and not kinetic considerations), they used the Wimley and White hydrophobicity scale, based on the free energy of transfer of amino acid side chains into nonpolar environments, to predict target peptide disposition with the membrane. The table below shows the propensity of amino acids to be in each region at equilibrium, based on this hydrophobicity scale.
Table: Amino Acid Partitioning Into Membrane Regions
| Region | Amino Acids |
| Bulk water | Arg, Asn, Asp, Gln, Glu, His, Lys, Pro |
| Bulk water + interfacial | Ala, Cys, Gly, Ser, Thr |
| Interfacial | Tyr |
| Hydrophobic | Ile, Leu, Met, Phe, Trp, Val |
Their experimental result were in concordance with those predicted using
the above scale. If a polyalanine 19 mer was used, no insertion was
observed. With five leucines in the peptide, almost 90% inserted into
the membrane. The results would be modeling using a two-state
equilibrium:
Peptide inserted <==> Peptide translocated.
They
then substituted each of the twenty amino acids into a given position into a
target peptide and used the results to develop an empirical scale for
membrane transfer, not one based on simple transfer to nonpolar medium.
This new scale matched the hydophobicity scale, suggesting insertion and
transfer decisions where based on thermodynamics of side chain partitioning.
They also varied the position of the varied amino acid in the test peptide.
If the amino acid favored the bulk and/or interfacial region, the peptide
would be inserted if that amino acid were at the end of the peptide, not the
middle. For translocation, the peptide had to be amphiphilic with one face
polar and the other nonpolar.
They developed a simple equilibrium model to show the processes involved, as shown below in a top-down view of the membrane.
Figure: Translocon Equilibrium Model
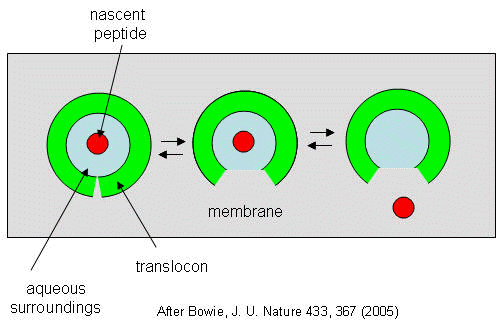
The translocon, shown in green, has a water-filled pore but also a sidewise opening toward the membrane interior. The target peptide enters the pore. Transient conformational changes in the pore expose the peptide to the nonpolar membrane core. The target peptide samples both the aqueous and nonpolar environments and partitions into them based on considerations mentioned above. If it partitions more favorably into the hydrophobic core, it will do so and cause the peptide to become membrane bound. Otherwise it will pass through to the other side. This can be modeled as an equilibrium process if the rate of translocation is slow compared to the rates of translocon conformational change and environmental sampling by the peptide. Obviously, the process becomes more complicated if the target is a large protein.
![]() Jmol:
Updated SecYEB Protein Translocase
Jmol14 (Java) |
JSMol (HTML5)
Jmol:
Updated SecYEB Protein Translocase
Jmol14 (Java) |
JSMol (HTML5)
Bacterial toxin proteins also have evolved ways to pass through a cell membrane, again in a nonnative state, through a protein channel in the membrane. Krantz et al have recently worked out details of how the anthrax toxin protein moves through eukaryotic cell membranes. Three anthrax proteins are involved. One is a "prepore" protein that binds to specific proteins on the cell membrane, where it is activated by limited proteolysis to form a pore protein which assembles into the homoheptamer prepore in the membrane. Two other proteins secreted by the bacteria, lethal factor and edema factor, bind to the heptamar complex and the whole assembly is then taken up into the cell by invagination to form a vesicle with the pore complex in the membrane. This vesicles fuses with a lysosome in the cell, and upon acidification, a conformational change occurs in the prepore complex to activate it. The lethal and edema factors unfold partially, possibly to a molten globule state, and are then passed though the pore into the cell where they exert their toxic influences. An electrochemical potential gradient (which we will discuss later in the semester) is required for passage of the factors through the membrane. The active pore further unravels the factor protein, facilitating transport.
Krantz et al. studied the pore protein by mutating two amino acids, Phe427 and Ser 429, on each monomer of the pore to Cys. They then postranslationally modified the Cys with [2-(trimethylammonium) ethylmethanethiosulfonate and observed effects on ion conductance of the pore and pore conformations. They noted that when both residues were mutated and chemically modified, that ion conductance was blocked, suggesting that these side chains were localized in the narrowest part of the channel. When Phe 427 alone was mutated to smaller side chains (Ala), ion conductance increased but transfer of peptides from the factor proteins was inhibited. This suggested that an aromatic ring in the narrow part of the channel opening participates in translocation of bacterial proteins through the membrane. They then analyzed the transport of a variety of small molecules with varying hydrophobicity through the wild type pore. Their results were consistent with the binding of the molecules through hydrophobic and aromatic electron interactions. They suggest a mechanism of transport consistent with their data in which the unfolded protein "ratchets" through the pore, which promotes factor protein unfolding to expose more hydrophobic groups to the nonpolar aromatic ring in the pore. This mechanism is similar to how the chaperone complex GroEL/GroES unfolds protein in its large central cavity, in a process which requires the chemical potential released by hydrolysis of ATP, not a transmembrane potential. In addition, the Sec61 translocon in the inner membrane of bacteria and in eukaryotic ER membranes also has a pore containing a ring of hydrophobic groups (Ile).
D9. Recent References
- F. Ulrich Hartl, Andreas Bracher & Manajit Hayer-Hartl. Molecular chaperones in protein folding and proteostasis. Nature 475, 324-332 (2011) doi:10.1038/nature10317
- Dunker, A et al. Intrinsically disordered proteins. Journal of Molecular Graphics and Modeling. 19, 26 (2001)
- Dunker, A. et al. The unfoldomics decade: an update on intrinsically disordered proteins.BMC Genomics 2008, 9(Suppl 2):S1 doi:10.1186/1471-2164-9-S2-S1
- Schuck, S. et al. J. Cell Biol 187, 525 (2009)
- Riemer, J. et al. Disulfide Formation in the ER and Mitochondria: Two Solutions to a Common Process. Science 324, 1284 (2009)
- Depuydt, M. et al. A Periplasmic Reducing System Protects Single Cysteine Residues from Oxidation. Science 326, 1109 (2009)
- Uversky, V & Dunker, A. Controlled Chaos. Science. 320, 1340 (2008)
- Bemporad, F. et al. Biological function in a non-native partially folded state of a protein. EMBO Journal 27, 1525 (2008)
- Murzin, A. Metamorphic Proteins. Science 320, 1725 (2008)
- Kimchi-Sarfaty, C. et al. A "Silent Polymorphism in the MDR1 Gene Alters Substrate Specificity. Science 315, 525 (2007)
- Religa, T. et al. Solution Structure of a protein denatured state and folding intermediate. Nature. 437, 1053 (2005)
- Cecconi et al. Direct Observation of the Three-State Folding of a Single Protein Molecule. Science 309, 2057 (2005)
- Krantz, B.A. et al. A Phenylalanine Clamp Catalyzes Protein Translocation Through the Anthrax Toxin Pore. Science 309, 777 (2005)
- Hessa, T. et al. Recognition of transmembrane helices by the endoplasmic reticulum translocon. Nature 433, 377 (2005)
- Bowie, J. U. Cell Biology: Border Crossing. Nature 433, 367 (2005)
- van den Berg, B et al. X-ray structure of a protein-conducting channel. Nature, 427, pg 36 (2004)
- Dobson, C. Protein folding and misfolding. Nature. 426, pg 884 (2003)
- Kamal, A. et al. A high-affinity conformation of Hsp90 confers tumour selectivity on Hsp90 inhibitors. Nature. 425, pg 407, 357 (2003)
- Ishii, D et al. Chaperonin-mediated stabilization and ATP-triggered release of semiconductor nanoparticles. Nature. 423, pg 628 (2003)
- Hartl and Hartl. Molecular Chaperones in the Cytosol: from nascent chain to folded protein Sicence. 295, og 1852 (2002)
- Houry et al. Identification of in vivo substrates of the chaperonin GroEL. Nature Nov 1999, pg 147. (Vol 402?)
- Vendruscolo et al. Three key residues form a critical contact network in a protein folding transition state. Nature 409, pg 641 (2001)
- Baker. A surprising simplicity to protein folding. Nature. 405, pg 39 (2000)
- Batey et al. Crystal structure of the ribonucleoprotein core of the signal recognition particle. (which binds newly synthesized proteins cotranslationally, arrests the synthesis, and docks the particle to the endoplasmic reticulum where synthesis restarts and the protein is discharged into the ER lumen for movement elsewhere in the cell). Science. 287, pg 1232 (2000)
- Shin et al. Interaction of partially unfolded forms of Torpedo acetylcholinesterase with liposomes. Protein Science. 5, pg 42 (1996)
- Ren et al. Interaction of diphtheria toxin T domain (transmembrane domain) with molten globule-like proteins and its implication for translocation. Science. 284. pg 955 (1999)
- Netzer and Hartl. Recombination of protein domains facilitated by co-translational folding in eukaryotes. Nature. 388, pg 329, 343 (1997)
- Weber-Ban et al. Global unfolding of a substrate protein (GFP) by the Hsp 100 chaperone ClpA. Nature. 401, pg 29, 90 (1999)

Biochemistry Online by Henry Jakubowski is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.