Reactivity in Chemistry
Reduction Potentials of Metal Ions in Biology
MB5. Hard and Soft Acid and Base Considerations
There are a number of amino acid residues that commonly bind to metal ions in biology. Do certain metal ions have a preference for certain donor atoms? And do those donor atoms have an influence on the properties of the metal complexes that are formed?
The principle of hard and soft acids and bases (HSAB) is useful here. Remember, this is a classification system that is based on the amount of charge density on acids and bases. Small ions with tightly held charges are called "hard". Larger, more polarisable ions are called "soft". Typically, hard cations (hard acids) combine more readily with hard anions (hard bases). Soft bases combine readily with soft acids.
Problem MB5.1.
Which of the following cations would be considered hard?
a) Hg+ b) Ti4+ c) Ca2+ d) Cu+
Problem MB5.2.
Which of the following anions would be considered hard?
a) F- b) I- c) CH3S- d) HO-
Remember that in the case of cations, increasing positive charge makes the cation smaller, because the increasing positive charge draws the remaining electrons closer to the nucleus. That means that some metals could conceivably be thought of as soft in one oxidation state but hard in another. A more highly charged cation would be harder.
A few ions are placed in the following table for reference, with extra attention to cases that occur on biology.
Table MB5.1. A table of common hard and soft acids and bases.
| hard | borderline | soft | |
| acids | H+, Li+, Na+, K+, Mg2+, Ca2+, Co3+, Fe3+ | Fe2+, Co2+, Ni2+, Cu2+, Zn2+ | Cu+, Ag+, Au+, Hg+, Cd2+, Hg2+ |
| bases | F-, HO-, H2O, Cl-, RO-, O2-, CH3CO2-, RNH2, CO32-, NO3- | NO2-, Br-, N2, pyridine, imidazole | RSH, RS-, S2-, CN-, CO, I-, C2H4, H-, CH3- |
Notice that some ions are in between the hard and soft categories. These borderline acids bond most strongly with borderline bases.
Problem MB5.3.
Choose the best match for the following ions.
a) Co2+ with CO32-, NO2- or S2-
b) Mg2+ with imidazole, CN- or CH3CO2-
c) Cu+ with RS-, pyridine or H2O
d) Fe2+ with N2, I- or HO-
e) Zn2+ with O2-, imidazole or RSH
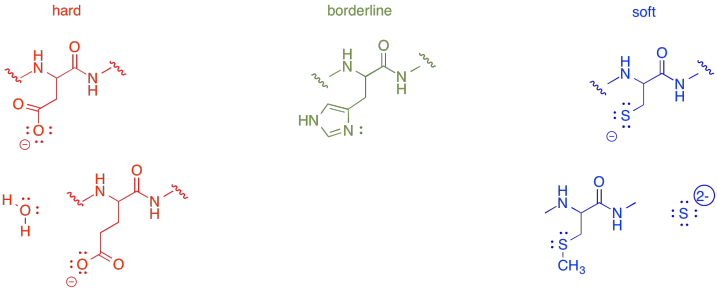
Many of the metal ions in the table above are biologically useful. From the information in the table, it shouldn't be too surprising that Fe2+ ions are often bound to histidine, which coordinates to the metal through an imidazole group. On the other hand, sometimes the same metals in different oxidation states may be coordinated by other amino acid residues. For example, higher oxidation state Fe3+ may more commonly have aspartate or glutamate donors.

Figure MB5.1. Some examples of hard and soft amino acid side chains.
Problem MB5.4.
Choose the best match for the following metal ions.
a) Fe3+ with asp, his or cys
b) Cu+ with glu, asp or met
c) Zn2+ with glu, his or cys
d) Cu2+ with his, H2O or S2-
e) Co3+ with glu, cys or met
It's important to get familiar with these relationships. However, metal ions are rarely coordinated only to one type of ligand in biology. There may be three or four different kinds of amino acids bound to the same metal ion in one complex. We don't just find the same metal ion bound to the same amino acid all the time.
Why is that? If Fe2+ forms the most stable complexes with histidine, why doesn't it just bind to histidine and nothing else? The answer is complicated, but has something to do with the fact that the metal ion will make do with the amino acids that are available at a specific site in the protein. Sometimes the match may not be ideal. That's to our advantage, though. After all, metal ions do not just play structural roles in biology. They are also supposed to do something.
Very often, a coordinated ligand can tune the reactivity of a metal ion. It does so not by stabilising the metal ion in its current state but by stabilising an alternative state, if only the metal ion would undergo a reaction to get there. A combination of amino acid residues and other ligands may act together to hold the metal ion in a fine balance between two oxidation states. After all, once a metal ion has reacted, it needs to get back to the way it was before. The cell can't afford to keep importing more trace elements just to keep going. It needs to recycle its reagents through catalysis.
Problem MB5.5.
Choose the amino acid residue that would have the effect on the metal ion as described, based on HSAB principles.
a) Increase the reduction potential of Cu2+: asp, his or met.
b) Decrease the reduction potential of Fe3+: glu, his or cys.
c) Make Cu+ easier to oxidise: his, cys or met.
d) Make Fe2+ easier to oxidise: asp, his or cys.
e) Make Fe3+ easier to reduce: asp, glu or cys.
This site is written and maintained by Chris P. Schaller, Ph.D., College of Saint Benedict / Saint John's University (retired) with contributions from other authors as noted. It is freely available for educational use.

Structure & Reactivity in Organic, Biological and Inorganic Chemistry by Chris Schaller is licensed under a Creative Commons Attribution-NonCommercial 3.0 Unported License.
This material is based upon work supported by the National Science Foundation under Grant No. 1043566.
Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the National Science Foundation.
Navigation: