Complex I: NADH-quinone oxidoreductase
Java version
HTML 5 version (does not require Java; downloads and moves
slowly)
I. Introduction
This is the most complicated membrane complex studied. We'll take it in
parts. First we'll look at the hyrophilic (matrix) domain where electron
transport from reduced NADH to ubiquinone occurs. Next we'll look at the
membrane domain where H+ transport occurs. Finally we'll look at the two
versions of the full complex. The pdb files are big so the Jmol files load
and move slowly. The figure below shows a carton
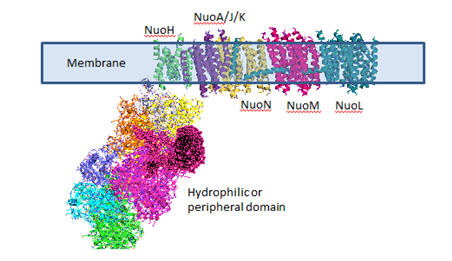
For more information see
Biochemistry Online: Chapter 2C - Understanding Protein
Conformation
II. General Structure
A. The hydrophilic domain - electron transport (pdb id: 3I9VA, THERMUS THERMOPHILUS, OXIDIZED)
Electron flow occurs from NADH to UQ through a series of one electron carriers in the hydrophilic or peripheral domain of complex 1. Initial handoff of electrons occurs to a flavin cofactor, FMN, and then through a series of Fe/S clusters. In the left figure below, these electron acceptors include a tetranuclear Fe/S cluster (SF4 shown yellow/red spacefill, a binuclear Fe/S cluster (FE2/S2) shown in blue, a FMN flavin
mononucleotide shown in CPK colors, .
Cartoon
add FMN - which receives electrons from NADH - Spacefill
add the tetranuclear FeS Clusters, Fe
4S
4 - Spacefill
add the 2 binuclear FeS Clusters, Fe
2S
2 - Spacefill
a single
tetranuclear FeS Cluster N3, Fe
4S
4, adjacent to FMN.
The cluster is based on the cubane structure with Fe and S occupying alternating corners of a square in a tetrahedral geometry. Each Fe is also coordinated to thiolate anions. The actual structure is a distorted cube
a single
binuclear FeS Cluster N1b, Fe
2S
2, adjacent to tetranuclear cluster N3
all tetra- and binuclear FeS Clusters with FMN without protein (binuclear clusters in blue spacefill)
H+ movement accompanying electron flow to the distal tetranuclear cluster, N2, probably involve the following side chaings: H169, H170, D86, R350, D401, H129, R279, H89, R125, E122, R249, Y257, Y254, Y260, R296
amino acids in D chain involved in H+ movement accompany electron transfer to N2)
B. The membrane domain - H+ transport (pdb id: 3RKO, THERMUS THERMOPHILUS)
Protons flow this transmembrane domain through a series of chains (NuoL, M, and N) that are
homologous in structure to proton/potassium ion antiporters
Cartoon;
Red (NuoL or B chain);
Yellow (NuoM or C chain);
Orange (NuoN or D chain);
Green (NuoA, J, and K
Note the helix in the plane of the membrane (left to right) that crosss across all the NuoL, M, and N chains. This probably, through a cantalever mechanism, allows concerted conformational changes in the H+ transporters.
C. The combined structure of Complex 1 from Thermus thermophilus (pdb id:3M9S from THERMUS THERMOPHILUS), missing some segments
Cartoon
D. The complete structure of eukaryotic (bovine) Complex I with associated proteins in the active state (pdb ID: 3LC5)
Cartoon
Note the additional protein changes involved in the complex.