Biochemistry Online: An Approach Based on Chemical Logic

CHAPTER 5 - BINDING
C: MODEL BINDING SYSTEMS
BIOCHEMISTRY - DR. JAKUBOWSKI
3/29/16
|
Learning Goals/Objectives for Chapter 5C: After class and this reading, students will be able to
|
C8. Binding, Intracellular Granules and Droplets
We've studied different types of protein aggregation, including aggregation of the native state (to form dimers, trimers, multimers, filaments) or alternate conformations (such as in prion protein aggregation and formation of inclusion bodies of misfolded proteins). It turns out that messenger RNA (which ultimately get translated into proteins) can aggregate with RNA binding proteins (RBPs) to form intracellular granules. These kinds of aggregates are commonly found in cell and have recently been recognized as non-membrane bound organelles. We've seen analogous particles, lipid droplets, which contain TAGs and cholesterol esters surrounded by a phospholipid monolayer with adsorbed protein, also promoted to the state of an organelle (in contrast to the recent demotion of the planet Pluto to a dwarf planet or plutoid). The lipid droplet however are surrounded by a "membrane", in this case a monolayer.
How do these and other granules form. A quick review of the Cell Tutorial (scroll to bottom) shows granule formation can be caused by a classic "phase transition", not unlike gaseous water can self associate through hydrogen bonds to form liquid drops which can freeze with the formation of more hydrogen bonds to form solids. Soluble biomolecules in cells can reversibly aggregate through the summation of multiple weak IMFs to form storage granules. This balance might be perturbed if storage granules aggregate further in a potentially irreversible process with health consequences. We've seen examples of the latter when neurodegenerative diseases like Alzheimer's and Mad Cow Disease. Lets delve into new insights into the processes involved in droplet formation.
![]()
Imagine small amounts of a sparing soluble oil added to an aqueous solution. Initially it is in solution, but at a higher concentration, London dispersion forces and the “hydrophobic effect” would drive the oil out of solution into liquid drops. This phase separation could also be called liquid-liquid demixing as two liquids (solubilized oil in water and separated oil drops) separate. This process has been shown to produce many types of non-membrane bound droplets (not to be confused with membrane bound vesicles) in the cell.
This phenomenon has also been seen with intrinsically disordered proteins. These are characterized by amorphous structures with repeated, often positively charged amino acids. Under the right condition, these can aggregate and “precipitate” from the solution. What is the nature of the precipitate? It might have properties more like distinct liquid droplets so this process could be called liquid-liquid demixing.
Properties of demixed drops would include reduced rates of diffusion of material into an out of the drop, coupled movements of materials in the drop, and probable weak hydrophobic-dependent aggregation making drops sensitive to agents like detergents. Liquid-like diffusion inside the drop is observed as evident by rapid recovery of fluorescence from partially photobleached internal components of the drop.
As with the formation of a crystalline solid from a liquid solution, the process must be seeded. For intrinsically disordered proteins, this process can be “catalyzed” by poly-(ADP-ribose), a nucleic acid-like polyanion. The negative charges would counter the positive charges in the disordered protein domain, which without neutralization, would interfere with protein/protein contacts necessary for aggregation/droplet formation and demixing. Aggregation in these cases may arise from hydrophobic interactions (even though hydrophobic side chains are underrepresented in the disordered domains).
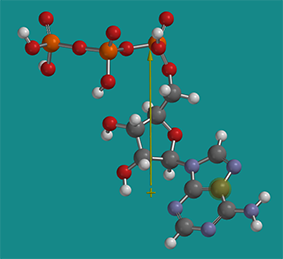
Solubility of proteins in cells is a fascinating topic in itself. It was recently discovered that the high concentration of ATP (5 mM) in the cell actually helps to solubilize proteins. ATP is considered a hydrotrope. It’s a small molecule with a vary distinct polar part (polyphosphate and ribose) and a more nonpolar part (the adenosine ring). Hence it acts sort of like a mini-detergent (an amphiphle) but it doesn’t form micelles. It does help stabilize more nonpolar parts of proteins in solution and has been shown to inhibit aggregate formation and also disaggregate some aggregates. The figure below shows a nonprotonated form of energy-minimized ATP with its dipole moment shown as an arrow from + to - end (obtained using Spartan). The dipole moment would only be larger if the ATP was deprotonated and had negative charges.
Biochemists also use the term gel (examples include polyacrylamide gel or fibrin blood clots which are chemically cross-linked) and a gel in the gel to liquid-crystalline phase transitions in lipid bilayers, held together by weak noncovalent interactions), when they wish to describe a structure that is neither clearly solid nor liquid. A structure like the cytoskeleton or the actin-myosin network would be examples of the latter.
Noncovalent gels would be characterized by regulatable dissociation of subunits and hence short half-lifes. A gel (either covalent or noncovalent) with a high-water content could be called a hydrogel which would contain hydrophilic components. An example would be RNA-protein containing particles
RNA granules
Many of the granules contain RNA and proteins and are called ribonucleoprotein bodies (RNPs) or RNA granules. Specific examples of these include cytoplasmic processing bodies, neuronal and germ granules, as well as nuclear Cajal bodies, nucleoli and nuclear dots/bodies). Some granules just contain proteins , including inclusion bodies with misfolded and aggregated proteins and those with active protein involved in biosynthesis, including the purinosome (for purine biosynthesis) and cellusomes (for cellulose degradation).
Another feature found in some neurodegenerative disease is a trinucleotide repeat. In Fragile X syndrome, there 230-4000 repeats of the CGG codon in the noncoding parts of the genome, compared to less than 50 in the normal gene. In Huntington’s disease, the repeat CAG is found in the protein coding part of the affected gene. The translated protein has a string of glutamines which probably causes protein aggregation. Specific proteins may also bind to the string of CAGs.
If the trinucleotide expansion is in intronic DNA, deleterious effects are not associated with translated proteins but with the transcribed RNA in the nucleus. The intronic repeats would be spliced out of the primary RNA transcript. A CTG DNA repeat would produce a poly CUG containing RNAs (found in myotonic dystrophy), which could aggregate through non-perfect base pairing. In vitro experiment show that small complexes are soluble, but as the size increases, a liquid-liquid demixing phase separation (or alternatively a liquid-gel transition) can occur, forming spherical drop RNA particles. This would explain the observation that pathologies occur above a certain repeat length. If misfolded proteins are also present, these particles might combine to form larger gels.
In the control experiment, when the repeats were scrambled, demixing and spherical particle formation was not observed. In an experiment similar to the addition of 1,6-hexanediol to intrinsically disorderd proteins, if small antisense trinucleotide repeats, such as (CTG)8, which could interfere with the weak H bonds between G and C in the aggregates, were added, the size of RNA drops (foci) were reduced. In vivo experiments showed characteristic drop-like structures but only if the repeats were of sufficient size.
Researchers found that in vitro, RNA drop formation was inhibited by monovalent cations. In the presence of 0.1 M ammonium acetate, which permeates cells without affecting pH, 47× CAG RNA droplets in vitro disappeared.
Aggregation of mRNA might be one way to regulate its translation and hence indirectly regulate gene activity. There are advantages to regulating the translation of a protein from mRNA, especially if the "activity" of the mRNA could be dynamically regulated. This would be useful if new protein synthesis was immediately required. Hence one way to regulate mRNA activity (other than degradation) is through reversible aggregation.
Protein drops and granules
The cytoskeletal proteins actin and tubulin (heterodimer of alpha and beta chains) can exist in soluble (by analogy to water gaseous) states or in condensed filamentous state (actin filaments and microtubules respectively). GTP hydrolysis is required for tubulin formation. Actin binds ATP which is necessary for filament formation but ATP cleavage is required for depolymerization. Hence nucleotide binding/hydrolysis regulates the filament equilibrium which differentiates from simple phases changes such as in water.
Since only certain proteins form granules, they must have similar structural features that facilitate reversible binding interactions. There appear to be multiple sites with these protein that individually form weak binding interactions, but collectively through multivalent (multiple) binding interactions allow robust but not irreversible granule formation. Here are some characteristics of proteins found in granules:
- the protein NCK has 3 repeated domains (SH3) the bind to proline-rich motifs (PRMs) in the protein NWASP. These proteins are involved in actin polymerizaiton. In high concentration they precipitate from solution and coalesce to form larger droplets;
- repeating interaction domains are widely found especially among RNA binding proteins;
- some proteins contain Phe-Gly (FG) repeats separated by hydrophilic amino acids in portions of the protein that are intrinsically disordered.
- a biotinylated derivative of 5-aryl-isoxazole-3-carboxyamide (structure below) precipitates proteins which are enriched in those that bind RNA (RBPs). In general the precipitate proteins were intrinsically disordered characterized by low complexity sequences (LCS). One such example contained 27 repeats of the tripeptide sequence (G/S)Y(G/S). The proteins could also form hydrogels (made of hydrophilic polymers and crosslinks) and transition between soluble and gel phases with extensive hydrogen bond networks. The hydrogel gel phase gave x-ray diffraction patterns similar to beta structure-enriched amyloid proteins. Short ranged weak interactions between LCS might then drive reversible condensation to gel like granule states characterized by extensive hydrogen bonding (again similar to hydrogen bonding on ice formation). If this process goes awry, more continued and irreversible formation of a solid fibril (as seen in neurodegenerative diseases) might occur from the hydrogel state;
-
RNAs appear in granules as protein bind them through RNA binding domains of proteins which interact through low complexity sequences leading to phase separation and hydrogel-like formation of granules. Around 500 RNA binding proteins have been found in the human RNA interactome. They are enriched in LCSs and have more tryosines than average proteins in the whole proteome in which the Tyr are often found in an (G/S)Y(G/S) motif. Phosphorylation of tyrosines (Y) in LCS may decrease association and hydrogel stability.
Given the many neurodegenerative diseases are associated with unfolded/misfolded protein aggregates, the high protein concentrations in protein-containing liquid drops might pose problems to cells. If high enough, the equilibrium might progress from the liquid drop to a solid precipitate, which would have severe cellular consequences. The progression to the solid state may irreversibly affect the cell.
In the next section, 5D: Binding and the Control of Gene Transcription, we will explore how liquid-liquid demixing can help explain chromatin structure and dynamics.
Navigation
Return to Chapter 5C: Model Binding Sections
Return to Biochemistry Online Table of Contents
Archived version of full Chapter 5C: Model Binding Systems

Biochemistry Online by Henry Jakubowski is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.