Biochemistry Online: An Approach Based on Chemical Logic

CHAPTER 8 - OXIDATION/PHOSPHORYLATION
B: OXIDATIVE ENZYMES
BIOCHEMISTRY - DR. JAKUBOWSKI
04/15/16
|
Learning Goals/Objectives for Chapter 8B: After class and this reading, students will be able to
|
B4. Another role of NAD - Sirtuins
Investigations of aging in many organisms have shown that a plethora of proteins seem to be involved. Studies have shown that overexpression of the yeast Sir2 (silence information regulator) caused an increase in lifespan. The enzyme product of this gene appear to be involved in inhibition, through covalent modification of chromatin and associated gene silencing, of genetic recombination. Investigators have shown that this protein is a NAD+-dependent histone deacetylase. Sir2 is a member of a large family of conserved protein deacetylases called sirtuins. Accompanying the deacetylase activity is the conversion of NAD+ to nicotinamide and acetyl-ADP ribose. Since the enzyme uses NAD+, its activity would be related to the metabolic state of the cell. It was thought that high levels of NAD+ would be present when metabolism was slow, and these high levels of NAD+, through its activation of Sir2, would lead to the silencing of genes and longer lifespan. Lack of genetic recombination would decrease DNA damage. Alternatively, low levels of nicotinamide (a product of Sir2) could minimize inhibition of Sir2 and raises its activity. However, Anderson et al. reported that in yeast, caloric restriction actually reduces NAD+, and that the activity of Sir2 does not depend on the ratio of NAD+/nicotinamide, suggesting that Sir2 is activated (during caloric restriction) by other molecules.
It has long been know that caloric restriction increases lifespans in a variety of animals. This provides a mechanism that would account for that. When glucose levels in culture media are restricted, the life span of the yeast increases, as measured by an increase in the reproductive life span of the organism. This effect is only observed if a functional Sir2 gene is present. Deacetylation of histone proteins by Sir2, would lead to an increased positive charge on the histones (protein components of the nucleosome), which presumably increases the binding affinity of histones (in nucleosomes) for DNA. This could reduce gene expression (i.e. induce silencing) since the nucleosomal DNA would be less available for transcription. Surprisingly, a precursor of cellular NAD+, nicotinamide, when given to cells to increase NAD+ levels which should silence gene expression had the opposite effect. The effect of Sir2 on life span has also been observed in a more complex organism, C. Elegans (round worm).
Another activity of Sir2 in yeast has been noted by Aguilaniu et al. When yeast cells divide, daughter cells do not inherit small DNA circles which code for ribosomal RNA. The concentration of these circles of DNA increase with age, and are associated with senescence of the maternal cells. Cells without Sir2 have more of these circles. Another feature of aging cells is the accumulation of oxidized proteins (as evidenced by an increase in carbonyl groups). In a fashion similar to rDNA circles, oxidatively-modified proteins are not inherited by daughter cells except in mutant yeast cells lacking Sir2.
Since increasing levels of Sir2 increase lifespan, ligands that bind to existing Sir2 and activate its deacetylase activity might also increase lifespan. Howitz et al. screened libraries of compounds for such ligands and found one exceptionally good one that activated the enzyme 13-fold: resveratrol, a polyphenol found in red wine.
Figure: resveratrol
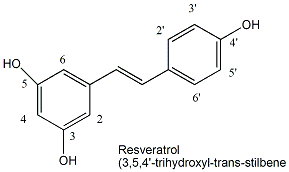
Studies have shown the people who drink moderate amounts of red wine may have lower risk for certain disease (for example some cancers and cardiovascular disease) and longer life spans. When given to yeast, resveratrol increased their life span by 70% in a process that required Sir2. Mutant strains lacking Sir 2 did not accrue the effect. Whether resveratrol levels found in wine are sufficient to promote this effect in humans is unclear. Wood et al. have recently extended the effects of resveratrol on activation of Sir2 and increased life-span to more complex organism - C. elegans and Drosophila melanogaster.
In humans there appears to be seven different sirtuins. Sirtuin 2 has been shown to deacetylate α-tubulin (involved in cycle cycle), p53 (a tumor suppressor gene) and histones H3 and H4. In some cases inhibition of sirtuin activity is beneficial.
The product of the gene DBC1 (deleted in breast cancer 1) has been shown to bind to and inhibit SIRT1, which causes increases in the concentration of acetylated p53 and other proteins regulated by it. One function of active p53 is to induce cell death (apotosis) in stressed cells (such as cancer cells). Hence sirutin 1 would inhibit p53 induced apotosis of cancer cells. .
Figure: Reaction of NAD+ and Acetyl-Lys side chains: possible catalytic mechanism for yeast Sir2
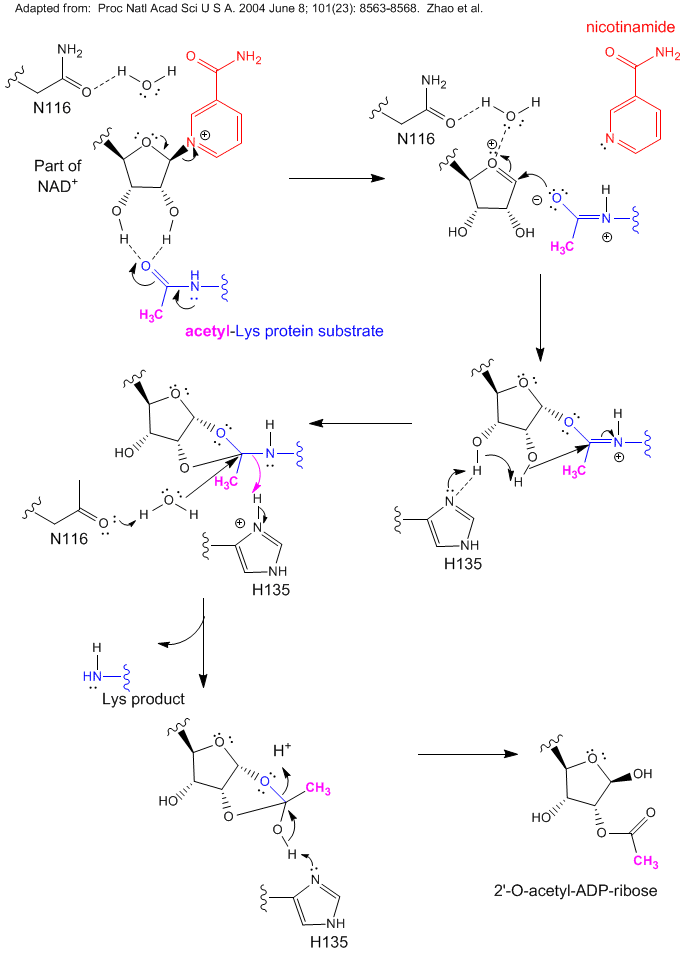
The effects of resveratrol has been extended to mammals through the study of its effects on mouse Sir1, one of seven similar proteins in mammals, is analogous to Sir2 in yeast. Sir1 is also a deacetylase which appears to work on many different acetylated proteins. It functions in part with another protein, peroxisome proliferator-activated receptor γ coactivator (PGC-1α), to regulate two important carbohydrate metabolism pathways in the liver under conditions of caloric restriction, probably through responses to NAD+ levels. The two pathways are glycolysis, the breakdown of the 6C glucose to the 3C molecule pyruvate, and gluconeogenesis, the synthesis of glucose from pyruvate. When blood sugar (glucose) is low, or in a diabetic state (when blood glucose levels are high due to impaired insulin function which facilitates glucose uptake into cells), sugar metabolism in the liver shifts to gluconeogenesis to produce glucose for export. This might sound contradictory in the diabetic state when blood glucose is abundant. However, the liver is responding to other signals in the blood sent from tissues which lack glucose, so its response is to synthesis glucose and release it into the blood. Sir1 deacetylates acetylated Lys side chains on PGC-1α, which increases its biological function and leads to transcription of liver genes for gluconeogenesis.
Two studies in mice extent the beneficial effects of resveratrol. In one study by Baur et al., mice fed a high calorie diet and who became obese avoided the negative effects of obesity if given moderate amounts of resveratrol. Compared to controls who were fed the same diet and not given resveratrol, the treated mice had a much longer life expectancy. Resveratrol appears to alter the deleterious changes in gene expressions in mice that are associated with insulin resistance, diabetes and cardiovascular disease. Resveratrol increased insulin sensitivity, decreased a hormone called insulin-like growth factor 1 (IGF-1, increased PGC-1αactivity, and mitochondrial numbers.
In another study by Lagouge et al, ordinary (untrained) mice given resveratrol had markedly increased endurance and oxygen consumption with decreased heart rate on a treadmill test, suggesting improved mitochondrial function. Resveratrol increased transcription of muscles (not liver) genes associated with mitochondrial oxidative phosphorylation (the topic of the next section) and with production of new mitochondria (mitochondria biogenesis). They studied mitochondrial function since this organelle is the site of most oxygen consumption and presumably the most reactive oxygen species (ROS), which have a role in aging and disease. Mitochondrial function also decreases with age. The effects of resveratrol on one particularly important gene involved in mitochondrial function, PGC-1α, was noted. Transcription of this gene in muscle is decreased in diabetics and in people resistant to insulin, both of which suffer impaired mitochondrial function. This protein has many biological functions, including increasing mitochondrial function and biogenesis. As noted above, resveratrol decreased acetylation of PGC-1α, and increased its function, which in muscle lead to increased transcription of genes involved in mitochondrial biogenesis an oxidative phosphorylation. They also showed that these effects were mediated through the Sir1 protein since PGC-1 acetylation was not affected by resveratrol in mutant mouse embryonic fibroblast cells with a nonfunctional SIR1 gene.
Navigation
Return to Chapter 8B: Oxidative Enzymes Sections
Return to Biochemistry Online Table of Contents
Archived version of full Chapter 8B: Oxidative Enzymes

Biochemistry Online by Henry Jakubowski is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.