Biochemistry Online: An Approach Based on Chemical Logic

CHAPTER 8 - OXIDATION/PHOSPHORYLATION
C: ATP AND OXIDATIVE PHOSPHORYLATION
BIOCHEMISTRY - DR. JAKUBOWSKI
04/15/16
|
Learning Goals/Objectives for Chapter 8C: After class and this reading, students will be able to
|
C5. Complex I - NADH-quinone oxidoreductase
Now lets explore electron transport in greater detail by looking at the mechanisms of two specific complex, I and IV. Before that look at a detailed view of the entry pathway of electron transport and oxidative phosphorylation.
Figure: Detailed View of Oxidative Phosphorylation (reprinted with permission from Kanehisa Laboratories and the KEGG project: www.kegg.org )
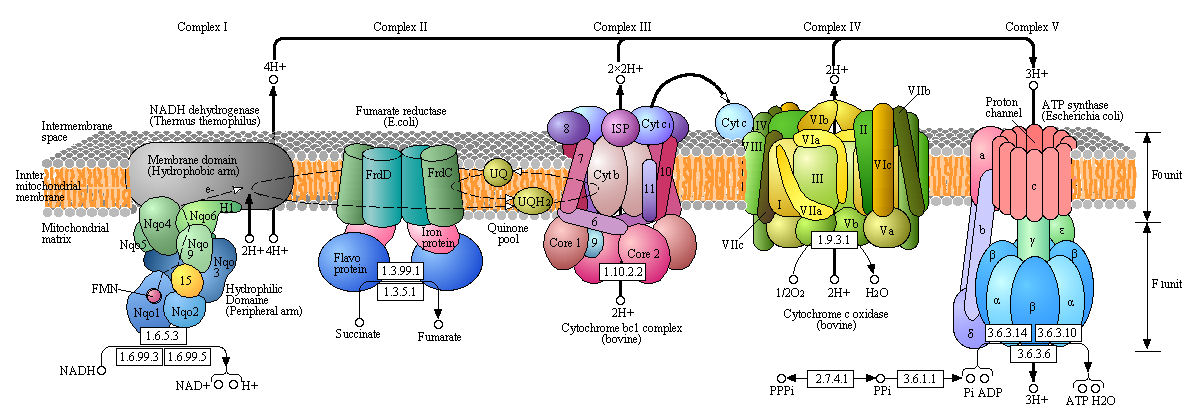
Boxed number represent Enzyme Commission Number. Original KEGG Map with imbedded links.
Complex I - NADH-quinone oxidoreductase
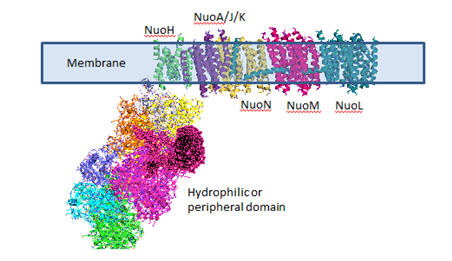
Complex I is located in the inner mitochondrial membrane in eukaryotes and in the plasma membrane of bacteria. Mammalian complex 1 consists of 45 subunits, 7 of which are encoded by mitochondrial gene. Bacteria have only 13-14 subunits. The significance of the extra mammalian subunits is still unclear. A cartoon model and the actual crystal structure of the Thermus thermophilus (bacterial) complex are shown below. The hydrophilic or peripheral domain catalyzes electrons transfer while the membrane domain (encoded by mitochondrial DNA) is involved in active transport of protons.
Figure: Detailed view of Complex I from Thermus Thermophilus
![]() 7/10/17: The following Jmol links contains multiple view of parts of
the complex as well as the entire complex. It is repeated several
times below.
7/10/17: The following Jmol links contains multiple view of parts of
the complex as well as the entire complex. It is repeated several
times below.
![]() Jmol:
Complex 1
Jmol14 (Java) |
JSMol (HTML5)
Jmol:
Complex 1
Jmol14 (Java) |
JSMol (HTML5)
Electron Transport
Electron flow occurs from NADH to UQ through a series of one electron carriers in the hydrophilic or peripheral domain of complex 1. Initial handoff of electrons occurs to a flavin cofactor, FMN, and then through a series of Fe/S clusters. In the left figure below, these electron acceptors include a tetranuclear Fe/S cluster (SF4 shown yellow/red spacefill, a binuclear Fe/S cluster (FE2/S2) shown in blue, a FMN flavin mononucleotide shown in red, and a MN (II) ion, shown in purple N1a and N1b are binuclear clusters and N2, N3, N4, N5, and N6 are tetranuclear clusters.
Figure: Electron Flow in Complex I from T. Thermophilus
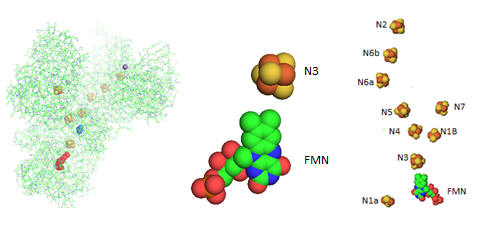
The tetranuclear Fe/S cluster is based on the cubane structure with Fe and S occupying alternating corners of a square in a tetrahedral geometry. Each Fe is also coordinated to thiolate anions. The actual structure is a distorted cube as shown below, along with that of the binuclear cluster, whose bond angle also deviate from those in a tetrahedron.
Figure: Fe/S Clusters in Complex I
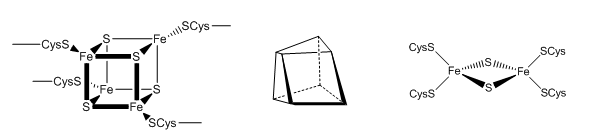
Many possible micro-redox states with different standard reduction potential are possible for tetranuclear Fe/S clusters, much as a polyprotic acid has multiple pKa values. The two relevant for Complex I and other tetranuclear clusters are shown below:
a. FeIIFe3IIIS4(CysS)41- + e- ↔ Fe2IIFe2IIIS4(CysS)42- (lower standard reduction potentials)
b. Fe2IIFe2IIIS4(CysS)42- + e- ↔ Fe3IIFeIIIS4(CysS)43- (higher standard reduction potentials)
Electrons are passed singly to oxidized UQ in one electron steps to form UQH2.
Fe-S clusters are synthesized predominately in the mitochondria where they serve as redox cofactors in electron transport as described above. They are ubiquitous in all life forms and serve roles in addition to redox cofactors per se as they serve structural roles in proteins and are used in redox signaling within the cell as they change oxidation states. Many proteins that interact with DNA (repair enzymes, polymerases and helicases) contain an Fe-S cluster.
Evidence suggests that they placed critical roles in the abiotic evolution of life in the absence of oxygen as a a terminal electron acceptor in exergonic oxidation reactions. When oxidation became available, they became potentially became toxic to the cell as the Fe2+ could participate in reactions (such as the Fenton reaction) leading to the generation of deleterious reactive oxygen species (such as superoxide). To prevent toxicity, when delivered to the cytoplasm and nucleus they must carried and delivered by cytoplasmic iron-sulfur assembly (CIA) proteins.
![]() Jmol:
Complex 1
Jmol14 (Java) |
JSMol (HTML5)
Jmol:
Complex 1
Jmol14 (Java) |
JSMol (HTML5)
Proton transport occurs in the membrane domain.
Available evidence suggests that 4 protons move from the cytoplasm to the periplasmic space against a concentration gradient during a catalytic cycle of Complex I in bacteria. One appears to be associated with the reduction of UQ at the terminal tetranuclear Fe/S cluster N2. The other three protons move across the membrane domain.
Nqo4 (proximal to the membrane domain as seen in KEGG diagram) residues in chain D have been implicated in H+ flow to the N2 cluster. These include, starting from the N2 cluster, H169, H170, D86, R350, D401, H129, R279, H89, R125, E122, R249, Y257, Y254, Y260, R296 (conserved residues are in bold). The terminal N2 cluster is coordinated by two tandem Cys side chains (C45 and C46) that in their thiolate (deprotonated form) are ligated to the Fe in the cluster. What properties do these amino acids have that make them candidates for this H+ flow?
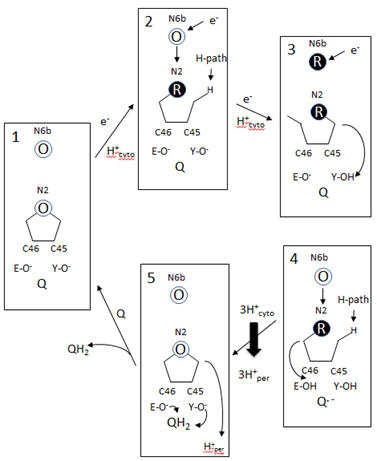
A model of electron and H+ flow is shown below (after Berrisford and Sazanov, JBC, 284, 29773, 2009). Iron-sulfur clusters N2 and N6b are depicted as O (for oxidized) or R (for reduced). C46 and C45 indicate the tandem cysteines from Nqo6 subunit (nearest the membrane domain). Q/QH2 indicate quinone/quinol. Tyr 87 (Y-O) and Glu 49 (D-O) are proton acceptors. H-path indicates proton delivery pathway from the cytosol to tandem cysteines. One proton from this cycle appears to be transported across the membrane. What happens to the other two protons shown in the diagram?
Figure: Coupled proton and electron transfer at the FeS center in Complex I
Additional proton are transported by the membrane domain. The NuoL, M, N, A/J/K and H transmembrane domains are shown below. The L, M and N domains have structures similar to proteins involve in the coupled movement of K+/H+ in opposite directions across a membrane (antiporter). There are discontinuous helices in each subunit. Possible residues at the discontinuity buried in the membrane helices are Glu 144 and Lys 234. Would you expect to find these buried in the membrane? If they are part of an unopened channel that is exposed on a concerted conformational change across 3 similar membrane protein domains, how would they participate in proton transfer.
Figure: Transmembrane Domain of Complex I of T. Thermophilus
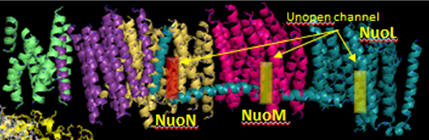
Looking at the left three antiporter subunits L, M, and N, notice a large helix that runs horizontally across all of them. How might that helix function to couple movement of protons across all the antiporter subunits (L, M, and N)? (Hint: think mechanically)
![]() Jmol:
Complex 1
Jmol14 (Java) |
JSMol (HTML5)
Jmol:
Complex 1
Jmol14 (Java) |
JSMol (HTML5)
Inhibition of Complex I
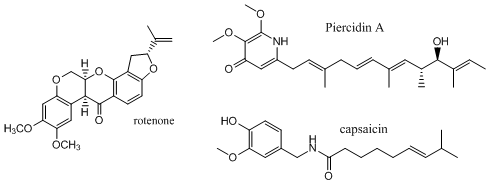
Complex I is inhibited by more than 60 different families of compounds. They include the classic Complex I inhibitor rotenone and many other synthetic insecticides/acaricides. The classes include: Class I/A (the prototype of which is Piericidin A), Class II/B (the prototype of which is Rotenone) and Class C (the prototype of which is Capsaicin). They appear to bind at the same site. From the structure of the 3 prototypes, what are the characteristics of the pharmacophore, the “ideal binding ligand”? Where do they likely bind? How “promiscuous” is the binding site?
Figure: Inhibitors of Complex I
Many devastating neurological diseases are associated with defects in Complex I. In addition to major problems with oxidative ATP production, reactive oxygen species (ROS) increase. The major sites for generation of ROS are Complex 1 and Complex III. Given the locations of the electron carriers at the periphery and internal within the protein complex, which electron carriers might most readily leak electrons to dioxygen? What ROS is likely to form in the process?
Inhibitors might block access of UQ or conformational changes necessary for final reduction of the ubiqinone free radical. Class A inhibitors dramatically increase ROS production. The actual site of ROS production in Complex is a bit controversial. One possible electron donor to dioxygen is FMN. Why is this a likely candidate? Mutants that lack N2 iron-sulfur cluster showed ROS production. Is this consistent with FMN site involvement in ROS production?
In submitochondrial preparations, normal Complex I activity occurs (which leads to formation of a sustained proton gradient). Also reverse electron transport, powered by an artificial proton gradient can occur, which leads to the reduction of NAD+ can occur (see diagram below).
Figure: Normal and Reverse Electron Transport Complex I
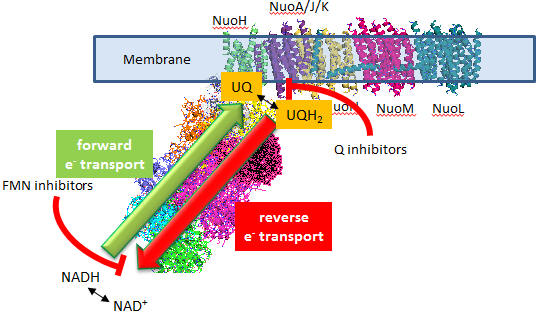
A summary of finding on superoxide production by Complex I is given below:
- Superoxide production is inhibited flavin site inhibitors but not Q site inhibitors.
- Reverse electron transport leads to NAD+ and O2 reduction
- Reverse electron transport superoxide production is inhibited by both flavin and Q site inhibitors
Based on these findings, which site, the flavin or Q site, is involved in superoxide production?
Navigation
Return to Chapter 8C: ATP and Oxidative Phosphorylation
Return to Biochemistry Online Table of Contents
Archived version of full Chapter 8C: ATP and Oxidative Phosphorylation

Biochemistry Online by Henry Jakubowski is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.