Biochemistry Online: An Approach Based on Chemical Logic

CHAPTER 8 - OXIDATION/PHOSPHORYLATION
C: ATP AND OXIDATIVE PHOSPHORYLATION
BIOCHEMISTRY - DR. JAKUBOWSKI
04/15/16
|
Learning Goals/Objectives for Chapter 8C: After class and this reading, students will be able to
|
C7. Complex IV - Cytochrome C oxidase (CCOx)
The structure of complex IV is shown in the left figure and to the right in a diagram taken from the KEGG pathways (with permission).
Figure: Cytochrome C Oxidase
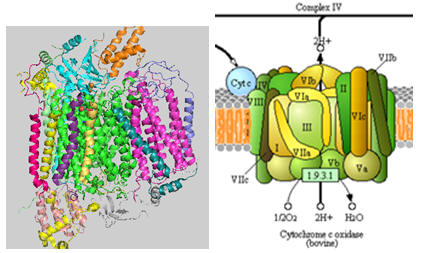
Cytochrome C, the initial “substrate” of this complex, delivers electrons from its heme cofactor to a dinuclear copper cluster, CuA. From there electrons flow to an adjacent heme a (low spin) which transfers them to another heme a3 (high spin) and then finally to dioxygen which is coordinated to the Fe in heme a3 and to an adjacent CuB. The heme a3 Fe:Cu dinuclear cluster is unique among all hemes. Which of the hemes is mostly likely to have two His side chains coordinated to the iron heme? One?
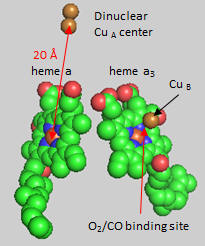
Figure: Heme and Fe:Cu cluster in Cytochrome C Oxidase - 1
![]() 7/13/17: The following Jmol links contains multiple views of the complex. It is repeated several
times below.
7/13/17: The following Jmol links contains multiple views of the complex. It is repeated several
times below.
![]() Jmol:
Complex IV
Jmol14 (Java) |
JSMol (HTML5)
Jmol:
Complex IV
Jmol14 (Java) |
JSMol (HTML5)
First let’s consider the transfer of electrons from heme a to heme a3 to dioxygen (we will consider entry of electrons into the complex later). What would be the consequence if dioxygen, a substrate for the reaction, dissociated from the heme a3 Fe before it were completely reduced? Suggest a reason for evolution of this key enzyme to have produced the unique heme a3 Fe:Cu dinuclear cluster. Heme a and a3 vary from the heme in hemoglobin as they both have a formyl group replacing a methyl and a hydroxyethylfarnesyl group added to a vinyl substituent. Its structure is shown below. What is its overall charge of the heme in its reduced state? In its oxidized state?
The key challenge has been to understand the redox coupling to H+ transport. How is this done? The formyl group on the heme is coplanar with the heme in both oxidation states. How might this coplanarity effect the charge in the heme?
Figure: Heme-Formyl group of Cytochrome C Oxidase
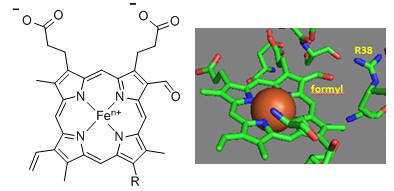
From the figure above, what type of interaction would likely occur between Arg 38 (R38) and the formyl group? What might occur to the protonation state of adjacent protein side chains, specifically Arg 38 (R38) on reduction of the heme? How would this link electron and proton transfer?
How might an electron be “transported” the 20 � distance from the dinuclear CuA cluster to the Fe in heme a and ultimately on to dioxygen? H+ transfer does not occur by physical movement of an individual proton through a “proton pore”. What is the most likely mechanism for transfer? Consider the ratio of the mass of a proton and an electron. Which is more likely to display wavelike behavior?
Crystal structures of oxidized and reduced CCOx show water channels and small “cavities” which calculations show can hold 1-3 water molecules. Hence groups around the heme, including R38 are assessable to water. What might be the function/role of the waters in the channel? Some of the amino acid residues associated with the water channels are shown in the figure below and include R38, S34, T 424, S461, S382, H413. How might these amino acids be involved in proton transfer? Draw an arrow on the diagram showing the direction of proton flow.
Figure: Proton Transfer in Cytochrome C Oxidase
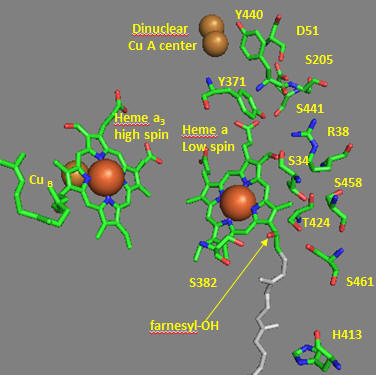
Another consequence of electron transfer to heme a involves the interaction of S382 and the farnesyl OH group, which are close in space and proximal to a water cavity. How might they interact? On reduction of the heme, a conformation change occurs which increases the S382-farnesyl OH group. What effect would this have on the interaction of the two and the S382-L381-Val380 localized conformation? A new water cavity appears to emerge on reduction in this region. How might this impact proton transfer from the matrix?
Now let’s consider the entry site of electrons into the complex and how they might influence proton transport. Structural and functional studies show a key role for Asp 51 (D51) (see figure above). In the oxidized state, D51 interactions with two OH side chains and amide NH backbone groups but is not exposed to water. On reduction, D51 lies on the surface in an aqueous environment. Draw both side chains in their likely protonation state in both the oxidized and reduced complex. What are the consequence of these structures for proton transport?
Near D51 is Y440-S441. The backbone carbonyl group between 440 and 441 forms an “indirect” interaction with R38 which we showed earlier is affected by the redox state of heme a. They are too far apart to form H bonds. Show by adding two waters how an interaction could occur between the carbonyl O and the side chain R38 via Y371. Also show how the water that interacts with Y371 also forms a H bond with the heme a proprionate.
Figure: Role of Amino Acids near D51 in Cytochrome C Oxidase
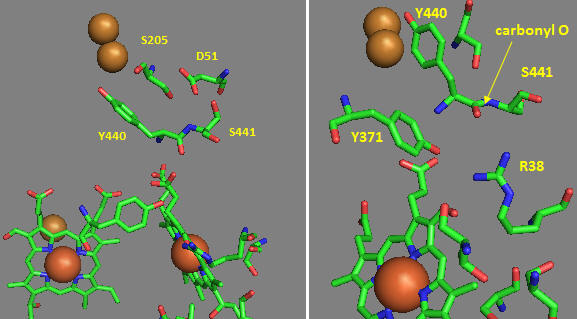
How could these groups participate in a proton transfer mechanism coupled to electron transfer? Would you expect a backbone carbonyl to be involved in proton transfer? Draw a reaction mechanism which shows how protonation of the carbonyl O could lead to formation of an imidic acid which leads to proton transfer from the backbone N. Also show reformation of the normal amide link through a tautomerization reaction. Why would this occur?
How does D51 connect to the this H bond network? In the oxidized state D51 is exposed on the surface. How might it change on reduction? Site specific mutagenesis has been used to change D51. What amino acid replacement might be optimal to affect activity but not protein folding? The mutation uncouples electron and proton transport. Which is likely to be affected?
A final pathway for coupling electron and proton transport has been proposed. Use the information above to complete the following statements: When heme a oxidized, Arg-38 is mainly ____________ (protonated/deprotonated) since _______ is available from the matrix. Asp 51 is ______________ (buried or exposed) and is ______________ (protonated/deprotonated). On reduction of heme a the net charge on heme a _________________ This leads to _________ (increased exposure/decreased exposure) of Asp-51 to the ___________ (intermembrane space, matrix, membrane) and _______ increased/decreased size of the water channel. Hence water molecules are _____________ (taken up/released from) the matrix. Coupled to this, protons on Asp-51 are ___________ (released to or taken up from) the intermembrane space On reoxidaiton of heme a, Asp-51 moves back to the ___________ (interior/exterior) of the protein and the net positive charge on heme a ___________ (increases or decreases) This leads to a _________ (increased or decreased) affinity of the heme formyl group for Arg 38. This leads to proton transfer __________ (to/from) Arg 38 toward Asp 51. Arg-38 then _____________ (gives up/take on) protons from water molecules in the water channel.
![]() 7/19/17
Mechanisms from higher resolution structures
7/19/17
Mechanisms from higher resolution structures
A more nuanced understanding of the mechanism and linkage between H+ and e- movement derives from high resolution structures determined by Yano et al (2016). In their model (shown in the figures below based on the oxidized form of the protein, pdb 5b1a), protons from the negative (matrix) N side of the complex enter through a water channel and proceed to the positve (intermembrane side) through a H bond network (as described above and depicted below). These comprise the H Pathway.
Directional movement is mediated by proton:proton repulsion aided by an increase in + charge on heme a when it transfers an electron to heme a3. Of course, proton:proton repulsion would move protons in both directions. Reverse flow back through the water channel is prevented by a conformational change on oxygen binding that closes the channel.
Ultimately 4 electrons are transferred from cytochrome Cs (in single electron steps) to the dicopper cluster, CuA, and then sequentially to heme a to heme a3 (near the copper B ion) to dioxygen to form water. The motion of electrons and protons are coupled electrostatically.
The figure below gives an overview of these movements. The small red dots are the oxygen atoms of internal water molecules (the rest have rest have been removed using Pymol). It should be apparent, given the number and location of the internal water molecules, that many would be involved in the proton translocation pathways.
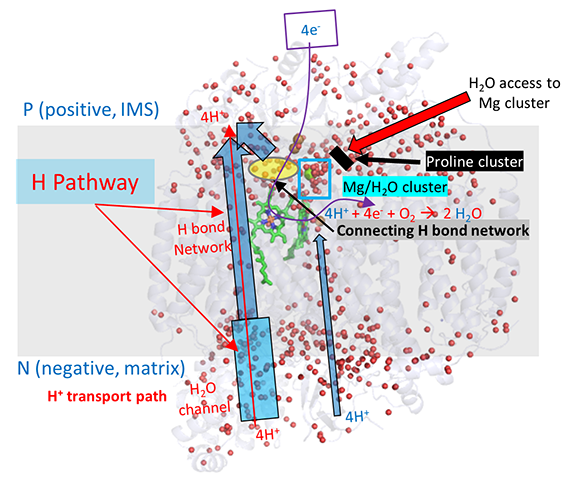
![]() Jmol:
Complex IV
Jmol14 (Java) |
JSMol (HTML5)
Jmol:
Complex IV
Jmol14 (Java) |
JSMol (HTML5)
What's so interesting about this model is the detailed description of two types of protons, the ones that add to dioxygen and end up in water, and those that are vectorially transported to the IMS. In their model, the H+s that end up being transported move through the water and H bond network through a connecting H bond link region to a Mg2+/water cluster. Since binding of oxygen leads to structural changes that closes off the water channel, all protons to be transported to the IMS must be bound in the cluster before dioxygen binding.
The figure below show that initially, 4 H+s move through the H system to the Mg2+/H2O cluster. Oxygen binding the closes the water channels. This buildup of positive charges would certainly lead to a enhanced electrostatic attractions for the next phase of the reaction, the movement of electrons into the heme cofactors. Additionally, the 4 H+s in the cluster are probably prevented from leaking to the P side through water that are proximal (see above figure) by proline cluster, which presumably restricts the dynamical motion of the protein in that region necessary for proton movement. The figure does not show charge changes in the electron carriers.
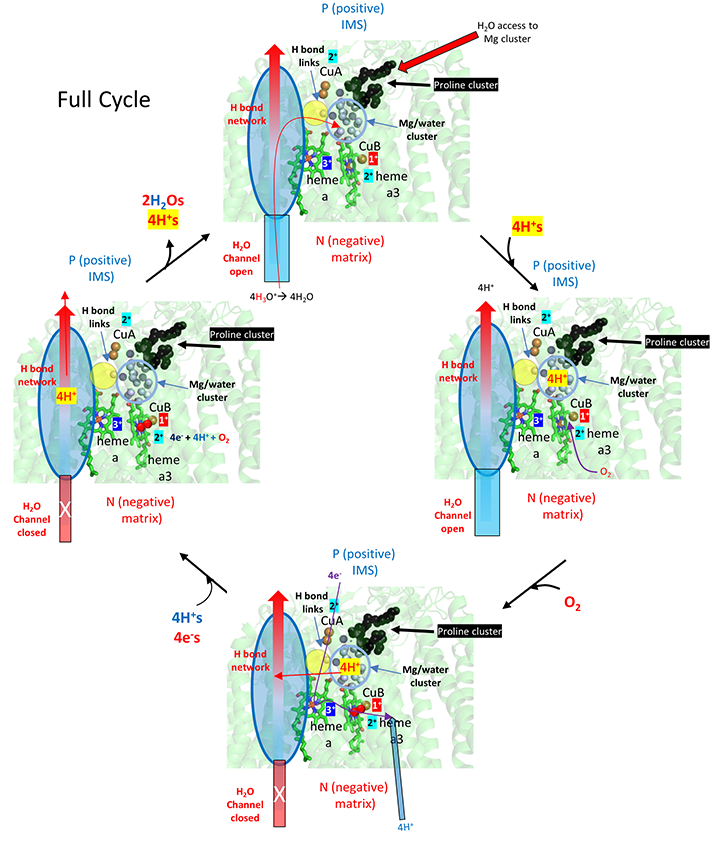
The figure below breaks downs the mechanism to show the addition of the first electron to the CuA (dicopper cluster), delivered from cytochrome C, and the subsequent transport of one proton from the fully proton-loaded Mg2+/water cluster after dioxygen binding. This figure does show the stepwise redox changes in the electron carriers.
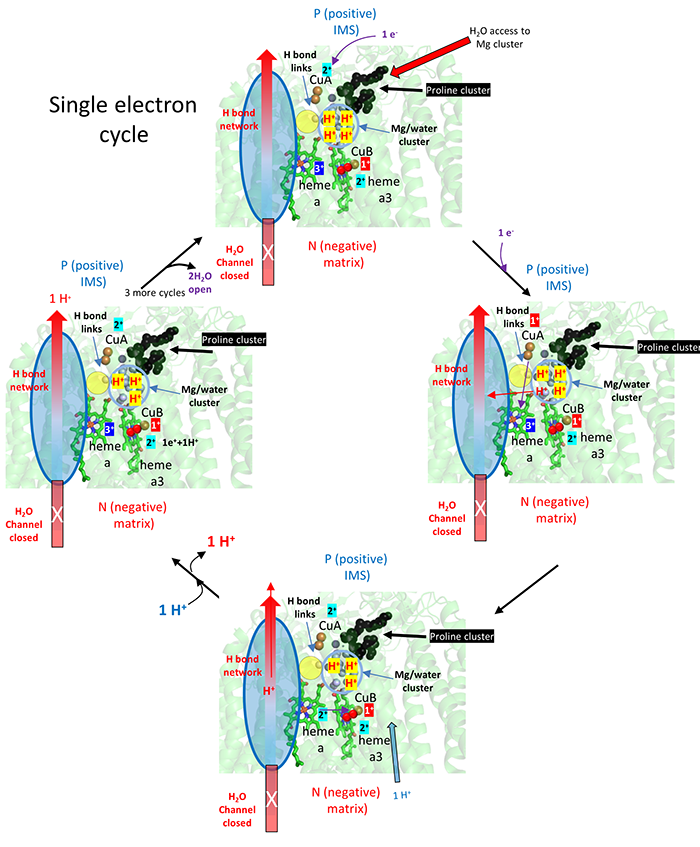
After CuA receives an electron from cytochrome C, it donates it to heme a and not to heme a3, even though both are close. The extra negative on heme a facilitates proton pumping thought the H pathways shown.
![]() Jmol:
Complex IV
Jmol14 (Java) |
JSMol (HTML5)
Jmol:
Complex IV
Jmol14 (Java) |
JSMol (HTML5)
Navigation
Return to Chapter 8C: ATP and Oxidative Phosphorylation
Return to Biochemistry Online Table of Contents
Archived version of full Chapter 8C: ATP and Oxidative Phosphorylation

Biochemistry Online by Henry Jakubowski is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.